import numpy as np
import pandas as pd
import plotnine as p9
import seaborn as sns
import scipy
import scipy.optimize
rng = np.random.default_rng(seed=123)Dimension reduction: t-SNE
t-SNE
t-SNE

t-SNE is a new-er-than-PCA method for dimension reduction (visualization of struture in high-dimensional data).
You can think about this as a lengthy exercise in “let’s dissect a sophisticated loss function for dimension reduction”.
It makes very nice visualizations.
The generic idea is to find a low dimensional representation (i.e., a picture) in which proximity of points reflects similarity of the corresponding (high dimensional) observations.
Suppose we have \(n\) data points, \(x_1, \ldots, x_n\) and each one has \(p\) variables: \(x_1 = (x_{11}, \ldots, x_{1p})\).
For each, we want to find a low-dimensional point \(y\): \[\begin{aligned} x_i \mapsto y_i \end{aligned}\] similarity of \(x_i\) and \(x_j\) is (somehow) reflected by proximity of \(y_i\) and \(y_j\)
We need to define:
- “low dimension” (usually, \(k=2\) or \(3\))
- “similarity” of \(x_i\) and \(x_j\)
- “reflected by proximity” of \(y_i\) and \(y_j\)
Similarity
To measure similarity, we just use (Euclidean) distance: \[\begin{aligned} d(x_i, x_j) = \sqrt{\sum_{\ell=1}^n (x_{i\ell} - x_{j\ell})^2} , \end{aligned}\] after first normalizing variables to vary on comparable scales.
Proximity
A natural choice is to require distances in the new space (\(d(y_i, y_j)\)) to be as close as possible to distances in the original space (\(d(x_i, x_j)\)).
That’s a pairwise condition. t-SNE instead tries to measure how faithfully relative distances are represented in each point’s neighborhood.
Neighborhoods
For each data point \(x_i\), define \[\begin{aligned} p_{ij} = \frac{ e^{-d(x_i, x_j)^2 / (2\sigma^2)} }{ \sum_{\ell \neq i} e^{-d(x_i, x_\ell)^2 / (2 \sigma^2)} } , \end{aligned}\] and \(p_{ii} = 0\).
This is the probability that point \(i\) would pick point \(j\) as a neighbor if these are chosen according to the Gaussian density centered at \(x_i\).
Exercise:
- Draw 9 points in a 10m x 10m square. Label the points 1 to 9.
- Draw another square, and try to draw the points 1 to 9 in that one so that the neighborhood of point
1is similar to the first picture, but the neighborhood of point9is not.
Reminder: the “neighborhood” is defined by relative distances: for point \(i\) it is \[\begin{aligned} p_{ij} = \frac{ e^{-d(x_i, x_j)^2 / (2\sigma^2)} }{ \sum_{\ell \neq i} e^{-d(x_i, x_\ell)^2 / (2 \sigma^2)} } , \end{aligned}\] and \(p_{ii} = 0\).
Similarly, in the output space, define \[\begin{aligned} q_{ij} = \frac{ (1 + d(y_i, y_j)^2)^{-1} }{ \sum_{\ell \neq i} (1 + d(y_i, y_\ell)^2)^{-1} } \end{aligned}\] and \(q_{ii} = 0\).
This is the probability that point \(i\) would pick point \(j\) as a neighbor if these are chosen according to the Cauchy centered at \(x_i\).
Similiarity of neighborhoods
We want neighborhoods to look similar, i.e., choose \(y\) so that \(q\) looks like \(p\).
To do this, we minimize \[\begin{aligned} \text{KL}(p \;|\; q) &= \sum_{i} \sum_{j} p_{ij} \log\left(\frac{p_{ij}}{q_{ij}}\right) . \end{aligned}\]
What’s KL()?
Kullback-Leibler divergence
\[\begin{aligned} \text{KL}(p \;|\; q) &= \sum_{i} \sum_{j} p_{ij} \log\left(\frac{p_{ij}}{q_{ij}}\right) . \end{aligned}\]
This is the average log-likelihood ratio between \(p\) and \(q\) for a single observation drawn from \(p\).
It is a measure of how “surprised” you would be by a sequence of samples from \(p\) that you think should be coming from \(q\).
Facts:
\(\text{KL}(p \;|\; q) \ge 0\)
\(\text{KL}(p \;|\; q) = 0\) only if \(p_i = q_i\) for all \(i\).
Summary (\(t\)-SNE):
Given data \(\{ x_i \in \mathbb{R}^n \}\), find locations \(\{ y_i \in \mathbb{R}^k \}\) to minimize \[\begin{aligned} \sum_{ij} p_{ij} \log(p_{ij} / q_{ij}) , \end{aligned}\] where \(p_{ij} \propto e^{-d_{ij}^2/\sigma^2}\) and \(q_{ij} \propto 1/(1 + d_{ij}^2)\), and rows in \(p\) and \(q\) are normalized to sum to 1.
Simulate data
set-up
A high-dimensional donut
Let’s first make some data. This will be some points distributed around a ellipse in \(n\) dimensions.
n = 20
npts = 1000
xy = rng.normal(size=n*npts).reshape((npts, n))
theta = rng.uniform(size=npts) * 2 * np.pi
theta.sort()
ab = rng.normal(size=2*n).reshape((2, n))
ab[:,1] = ab[:,1] - ab[:,0] * np.sum(ab[:,0] * ab[:,1]) / np.sqrt(np.sum(ab[:,1]**2))
ab /= np.sqrt(np.sum(ab**2, axis=1))[:,np.newaxis]
for k in range(npts):
dxy = 4 * np.array([np.cos(theta[k]), np.sin(theta[k])]).dot(ab)
xy[k,:] += dxyHere’s what the data look like:
sns.pairplot(pd.DataFrame(xy))
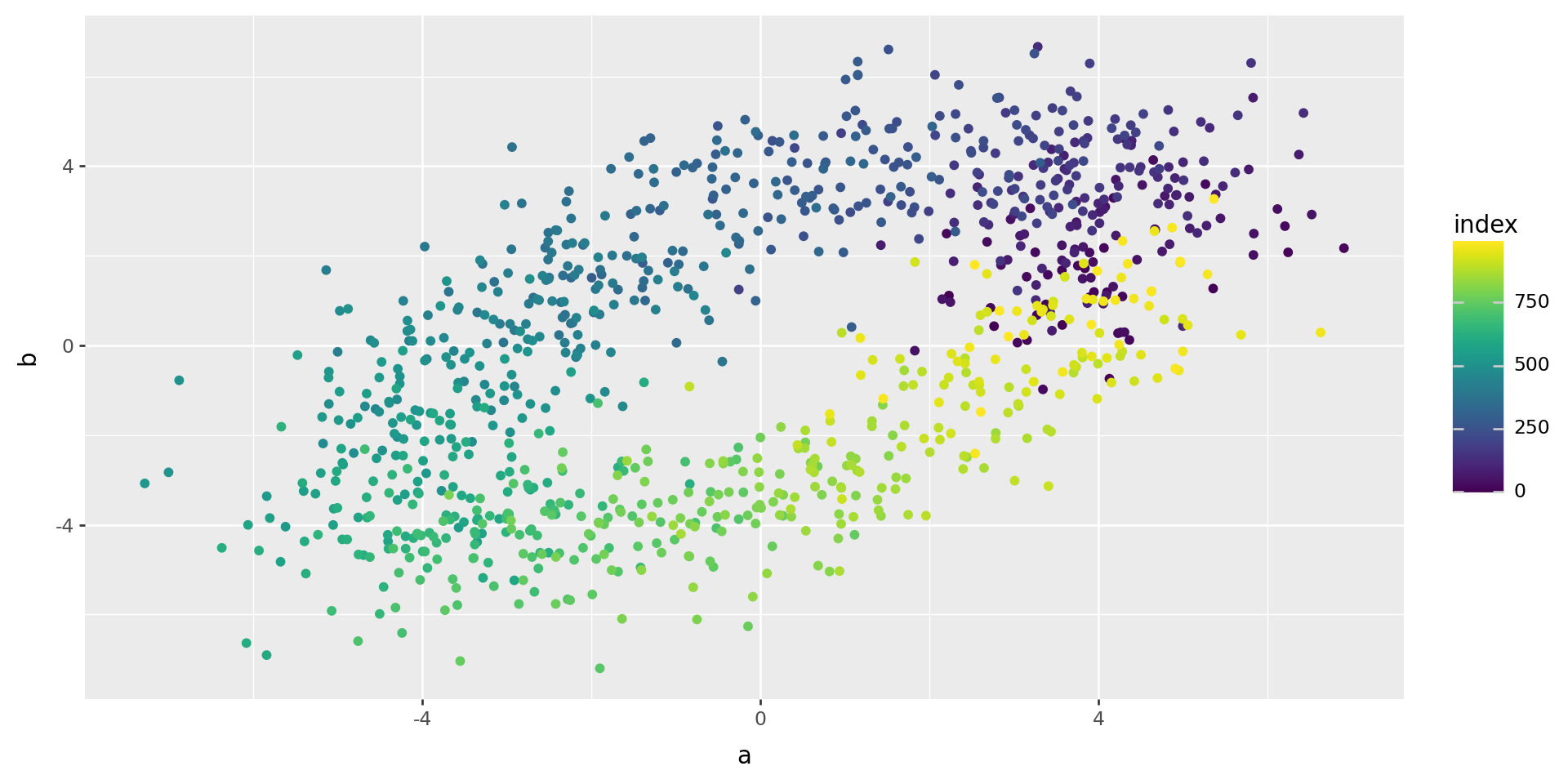
But there is hidden, two-dimensional structure:
(
pd.DataFrame(xy.dot(ab.T), columns=('a', 'b')).reset_index()
>>
p9.ggplot(p9.aes(x='a', y='b', color='index')) + p9.geom_point()
)
Okay, so again:
Given data \(\{ x_i \in \mathbb{R}^n \}\), find locations \(\{ y_i \in \mathbb{R}^k \}\) to minimize \[\begin{aligned} \sum_{ij} p_{ij} \log(p_{ij} / q_{ij}) , \end{aligned}\] where \(p_{ij} \propto e^{-d_{ij}^2/\sigma^2}\) and \(q_{ij} \propto 1/(1 + d_{ij}^2)\), and rows in \(p\) and \(q\) are normalized to sum to 1.
You could implement \(t\)-SNE, with that description.
But, let’s lean on skelearn instead:
import sklearn.manifold
tsne = sklearn.manifold.TSNE(n_components=2).fit_transform(xy)
tsnearray([[-15.271002 , 13.410212 ],
[ -7.4323015, 6.672564 ],
[ -9.591162 , 18.73543 ],
...,
[-19.885218 , 1.1804374],
[ -3.4351227, 16.46511 ],
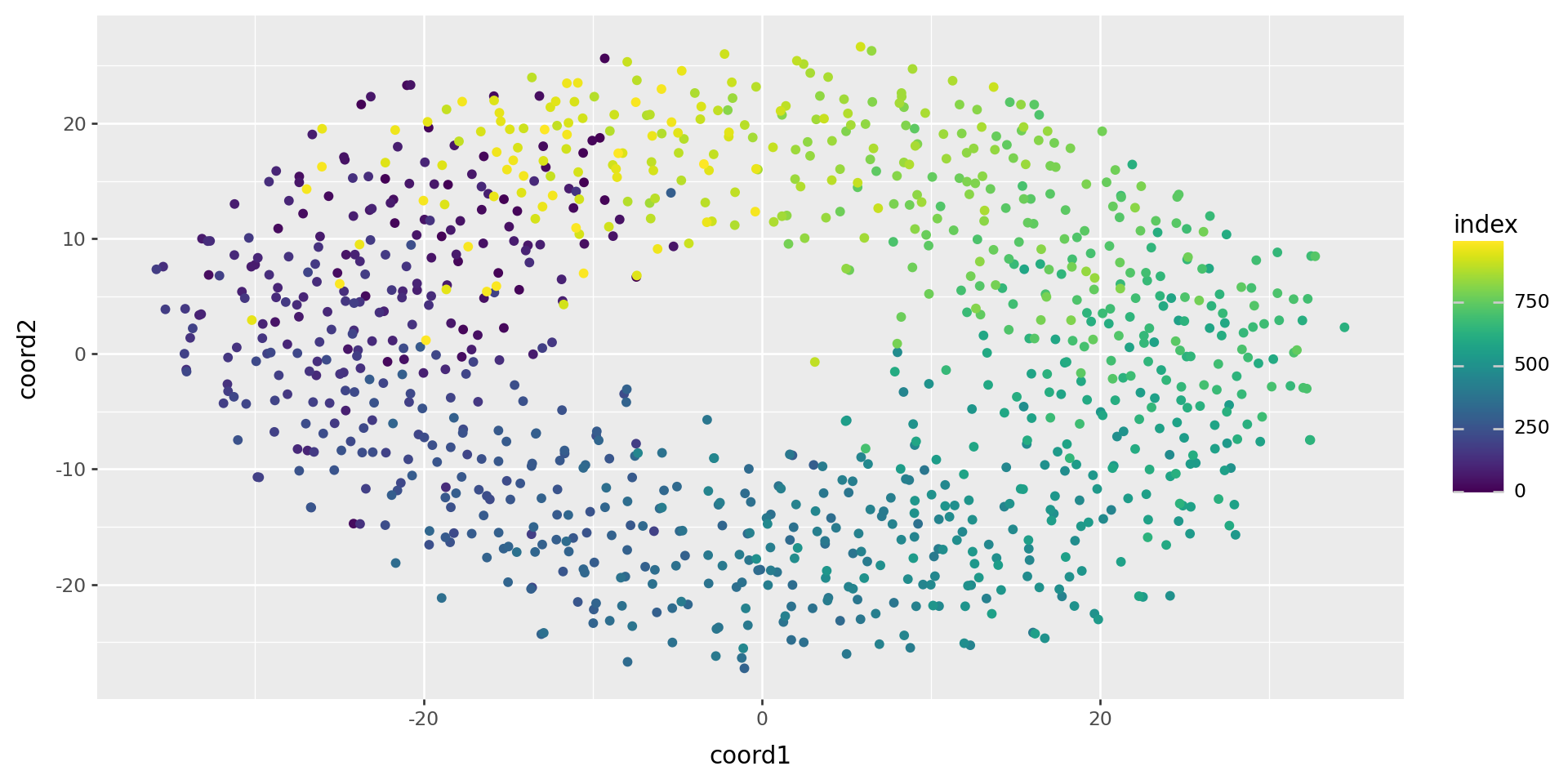
[-12.860488 , 19.459242 ]], shape=(1000, 2), dtype=float32)It works!
(
pd.DataFrame(tsne, columns=["coord1", "coord2"]).reset_index()
>>
p9.ggplot(p9.aes(x="coord1", y="coord2", color='index'))
+ p9.geom_point()
)
Another case
High-dimensional random walk
n = 40
npts = 100
rw = rng.normal(size=n*npts).reshape((npts, n))
for i in range(1, npts):
rw[i,:] = np.sqrt(.9) * rw[i-1,:] + np.sqrt(.1) * rw[i,:]sns.pairplot(pd.DataFrame(rw[:,:10]))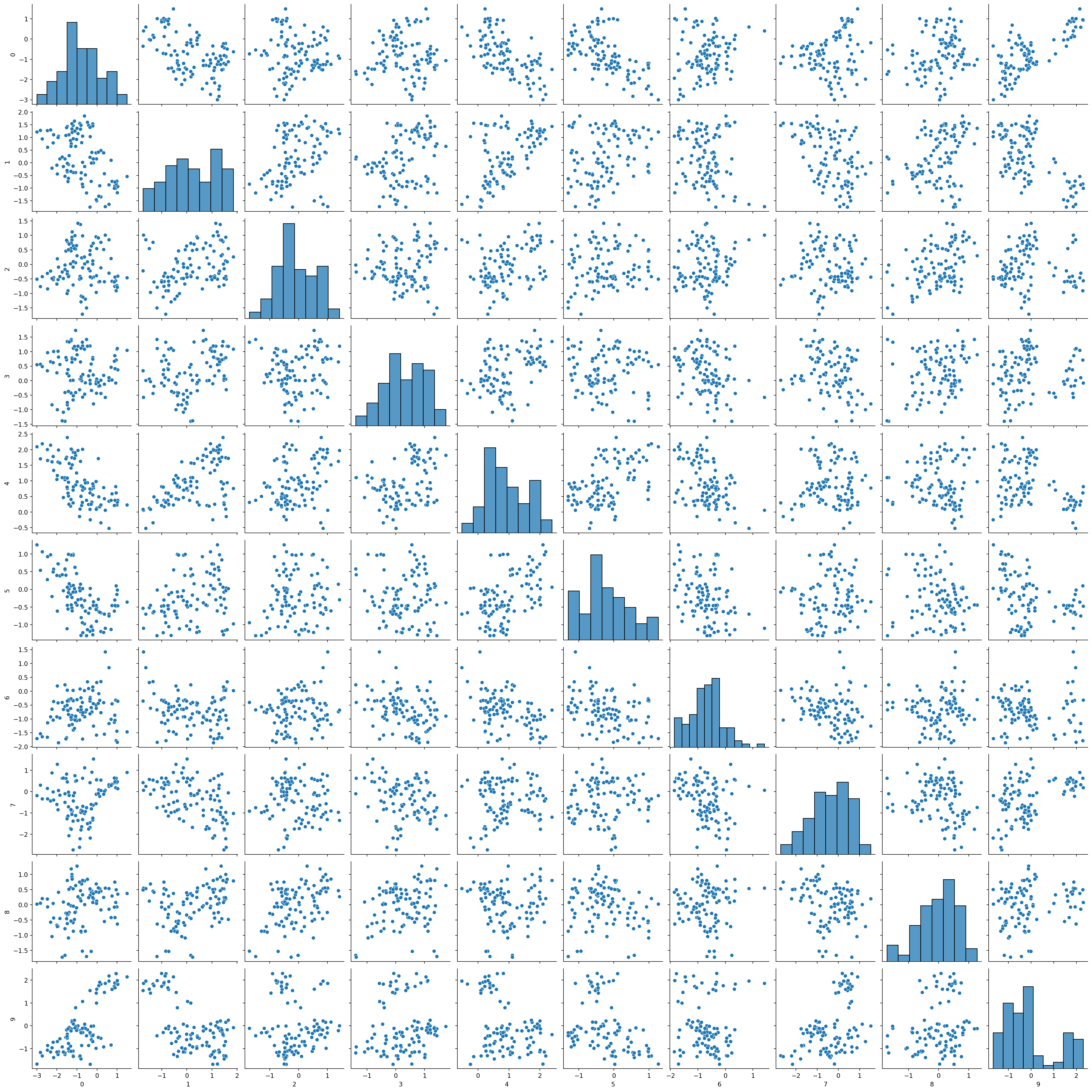
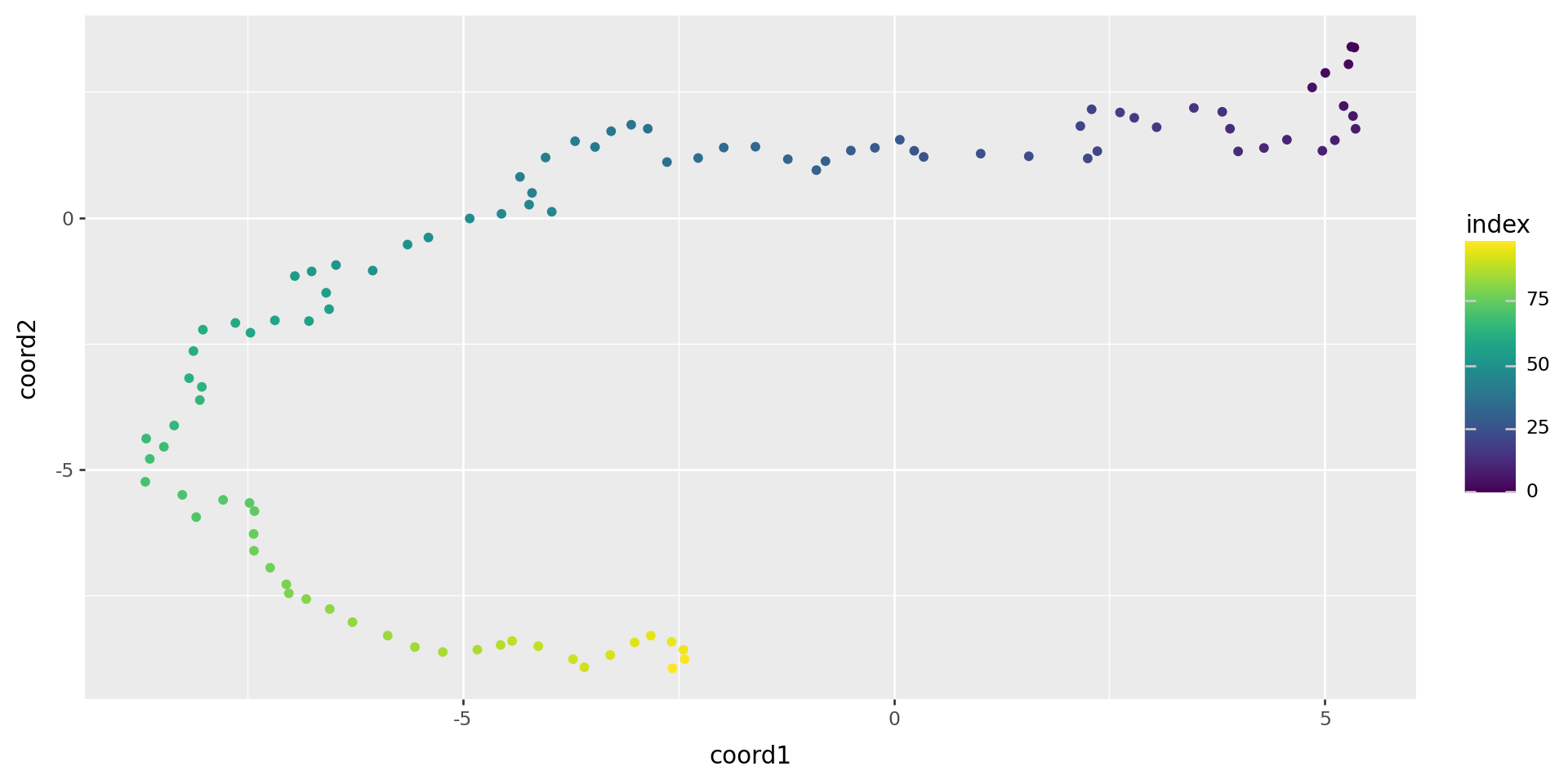
Run again
rw_tsne = sklearn.manifold.TSNE(n_components=2).fit_transform(rw)
rw_tsnearray([[ 5.342569 , 3.3803456 ],
[ 5.3078027 , 3.395333 ],
[ 5.274957 , 3.0481176 ],
[ 5.0061064 , 2.876894 ],
[ 4.8534102 , 2.5876067 ],
[ 5.2187524 , 2.218719 ],
[ 5.326692 , 2.0201669 ],
[ 5.35684 , 1.7662094 ],
[ 5.116625 , 1.5398095 ],
[ 4.971765 , 1.3300792 ],
[ 4.5605392 , 1.5498403 ],
[ 4.293864 , 1.3859835 ],
[ 3.9943395 , 1.317881 ],
[ 3.900134 , 1.7681946 ],
[ 3.8101308 , 2.1038268 ],
[ 3.4808762 , 2.1810582 ],
[ 3.048764 , 1.7975801 ],
[ 2.7894828 , 1.9848588 ],
[ 2.624019 , 2.0911021 ],
[ 2.2946653 , 2.153475 ],
[ 2.1637692 , 1.8207891 ],
[ 2.3605924 , 1.3225247 ],
[ 2.249378 , 1.1781802 ],
[ 1.5655448 , 1.2215273 ],
[ 1.0072862 , 1.2731194 ],
[ 0.34624624, 1.2084948 ],
[ 0.23634873, 1.3298091 ],
[ 0.06915712, 1.549084 ],
[-0.21990347, 1.3892608 ],
[-0.49869773, 1.334521 ],
[-0.7933359 , 1.1242819 ],
[-0.8989207 , 0.9460818 ],
[-1.2302965 , 1.163133 ],
[-1.6059532 , 1.4102476 ],
[-1.9735758 , 1.3939599 ],
[-2.2703006 , 1.1863585 ],
[-2.6321774 , 1.1061187 ],
[-2.8548625 , 1.7682289 ],
[-3.0470102 , 1.8471712 ],
[-3.279944 , 1.7185476 ],
[-3.4666314 , 1.4054004 ],
[-3.698554 , 1.519514 ],
[-4.040825 , 1.1959698 ],
[-4.3375406 , 0.8117907 ],
[-4.1977534 , 0.49342927],
[-4.2314615 , 0.2603935 ],
[-3.9689565 , 0.11954585],
[-4.5524664 , 0.07790056],
[-4.920539 , -0.01430212],
[-5.3997974 , -0.39193767],
[-5.641815 , -0.53180575],
[-6.0464735 , -1.0483378 ],
[-6.471822 , -0.93867815],
[-6.7545137 , -1.0654662 ],
[-6.9495997 , -1.157594 ],
[-6.586259 , -1.4898618 ],
[-6.5522013 , -1.8145885 ],
[-6.785826 , -2.0510683 ],
[-7.1808596 , -2.0374916 ],
[-7.4647293 , -2.2829072 ],
[-7.639564 , -2.0881324 ],
[-8.016611 , -2.2220306 ],
[-8.126271 , -2.646798 ],
[-8.176487 , -3.18546 ],
[-8.0286045 , -3.357377 ],
[-8.052256 , -3.6196513 ],
[-8.350244 , -4.124479 ],
[-8.674382 , -4.384362 ],
[-8.469013 , -4.5470223 ],
[-8.632379 , -4.7874694 ],
[-8.6844425 , -5.241904 ],
[-8.255149 , -5.50136 ],
[-8.094065 , -5.9435153 ],
[-7.781794 , -5.6013656 ],
[-7.475379 , -5.6623573 ],
[-7.4183664 , -5.824605 ],
[-7.428164 , -6.278005 ],
[-7.423861 , -6.6122 ],
[-7.235864 , -6.948705 ],
[-7.047818 , -7.278152 ],
[-7.019246 , -7.4556355 ],
[-6.817366 , -7.5708995 ],
[-6.544499 , -7.769315 ],
[-6.2812705 , -8.029693 ],
[-5.8719425 , -8.296413 ],
[-5.5569296 , -8.5241375 ],
[-5.2331185 , -8.621659 ],
[-4.8320847 , -8.576095 ],
[-4.562635 , -8.483491 ],
[-4.4285564 , -8.403036 ],
[-4.124957 , -8.507021 ],
[-3.722636 , -8.76384 ],
[-3.5903168 , -8.924602 ],
[-3.2904298 , -8.680731 ],
[-3.0074747 , -8.432109 ],
[-2.8198736 , -8.296976 ],
[-2.5770667 , -8.420178 ],
[-2.4419682 , -8.5795965 ],
[-2.4269888 , -8.764668 ],
[-2.5684159 , -8.945712 ]], dtype=float32)It works!
(
pd.DataFrame(rw_tsne, columns=["coord1", "coord2"]).reset_index()
>>
p9.ggplot(p9.aes(x="coord1", y="coord2", color='index'))
+ p9.geom_point()
)