Spatial smoothing
2026-02-25
Spatial point data: Biketown

Albert in a bike (catmapper:Wikimedia)

biketown bikes (Steve Morgan: Wikimedia)
how biketown works: screenshot
August, 2018
Here is data from August, 2018.
| RouteID | PaymentPlan | StartHub | StartLatitude | StartLongitude | StartDate | StartTime | EndHub | EndLatitude | EndLongitude | EndDate | EndTime | TripType | BikeID | BikeName | Distance_Miles | Duration | RentalAccessPath | MultipleRental | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | 8530077 | Casual | NW Couch at 11th | 45.523742 | -122.681813 | 8/1/2018 | 0:00 | NE Couch at MLK | 45.523588 | -122.661918 | 8/1/2018 | 0:09 | NaN | 6841 | 0075 BIKETOWN | 1.09 | 0:08:41 | mobile | False |
| 1 | 8530088 | Casual | NE Couch at MLK | 45.523588 | -122.661918 | 8/1/2018 | 0:01 | SW 2nd at Pine | 45.521421 | -122.672629 | 8/1/2018 | 0:08 | NaN | 7312 | 0987 BETRUE TRAINER | 0.62 | 0:07:24 | mobile | False |
| 2 | 8530156 | Subscriber | SW 3rd at Ankeny | 45.522479 | -122.673297 | 8/1/2018 | 0:07 | NaN | 45.522987 | -122.681129 | 8/1/2018 | 0:31 | NaN | 6145 | 0567 BIKETOWN | 2.62 | 0:23:36 | keypad | False |
| 3 | 8530170 | Subscriber | NaN | 45.520015 | -122.682115 | 8/1/2018 | 0:08 | NaN | 45.536330 | -122.685748 | 8/1/2018 | 0:18 | NaN | 7498 | 0181 BIKETOWN | 1.36 | 0:09:51 | keypad_rfid_card | False |
| 4 | 8530168 | Subscriber | NaN | 45.520015 | -122.682115 | 8/1/2018 | 0:08 | NaN | 45.536330 | -122.685748 | 8/1/2018 | 0:18 | NaN | 6879 | 0747 BIKETOWN | 1.36 | 0:09:39 | keypad_rfid_card | False |
Transform points
First, we’d like actual spatial points in a sensible (isometric) coordinate system:
df['start_points'] = gpd.points_from_xy(df.StartLongitude, df.StartLatitude, crs="EPSG:4326").to_crs("EPSG:5070")
df['end_points'] = gpd.points_from_xy(df.EndLongitude, df.EndLatitude, crs="EPSG:4326").to_crs("EPSG:5070")
gdf = gpd.GeoDataFrame(df, geometry=df['start_points'], crs=5070)
gdf[:1000].explore()Goals
- Where do most trips start?
- Where do they end up?
- Is there a net flux of bikes anywhere?
We’ll make some maps with spatial smoothing to answer this, and evaluate if that was a good tool for the job.
Interlude: when there’s lots of data
Too much data?
In this class we’ve mostly not worried about computation. Modern computers deal with vectors of length \(10^7\)–\(10^8\) pretty much instantly.
But, what if you’ve got more? Options:
- Use less data.
- Look at summaries of the data.
Considerations:
- Do you really need \(n=10^{12}\) for whatever you’re looking for? Probably not!
- If you sub-sample, make sure the sub-sampling is appropriate (i.e., if the sub-sample has structure, it’s structure you want to look at).
- You probably want to pilot your summaries on a sub-sample anyhow.
Too much data, take two
It’s real easy to run into “too much data” in spatial contexts, basically because \(N^2 > N\).
Why?
Images/rasters: my display is 2256 x 1504, which is 3.3M pixels. It’s easy to have a lot of images.
Pairwise comparisons: Many spatial operations require operating “nearby” objects. Naively, that requires working with all \(O(N^2)\) pairwise distances.
Example: kernel density estimation
Suppose we have a bunch of points
\[ (x_1, y_1), \ldots, (x_n, y_n) \]
with density \(f(x,y)\), i.e.,
\[ f(x,y) \approx \frac{\#(\text{of points within $r$ of $(x,y)$})}{\pi r^2} \]
Using (for instance)
\[ \rho_\sigma(x,y) = \frac{1}{2\pi\sigma^2} e^{-\frac{x^2+y^2}{2\sigma^2}} \]
we can estimate \(f(x,y)\) by
\[ \hat f(x,y) = \sum_{i=1}^n \rho_\sigma(x_i-x, y_i-y) . \]
(interlude: why?)
Try it out (in one dimension)
Simulate 200 spatial locations with a density having two “bumps”. Plot these points. (
rng.normal(size=n, loc=-3), rng.normal(size=n, loc=+3))Make 20 “reference” locations.
Compute the kernel density estimate for each reference location, with \(\sigma\) chosen appropriately, and plot it. (
scipy.stats.norm( ))
Uh-oh, it’s quadratic
Note that this required \(O(NM)\) operations, where \(N\) is the number of data points, and \(M\) is the number of reference locations.
This is not as bad as \(O(N^2)\)!
But, suppose we wanted “density estimate for each of our \(N\) points”. The naive method would use data points = reference points: \(O(N^2)\).
Better: get estimated density at a grid of references, and use linear interpolation.
Guidelines for keeping your computer happy
- Start with small \(N\), look out for things that take a long time.
- If you actually need “all pairwise” values, keep \(N\) small (below 1,000 or so?).
- If an operation seems to need pairwise values under the hood, use “algorithms”: for instance, \(k-d\) trees (spatial libraries will do this sort of thing for you).
Back to the data
Smoothed map of starting locations
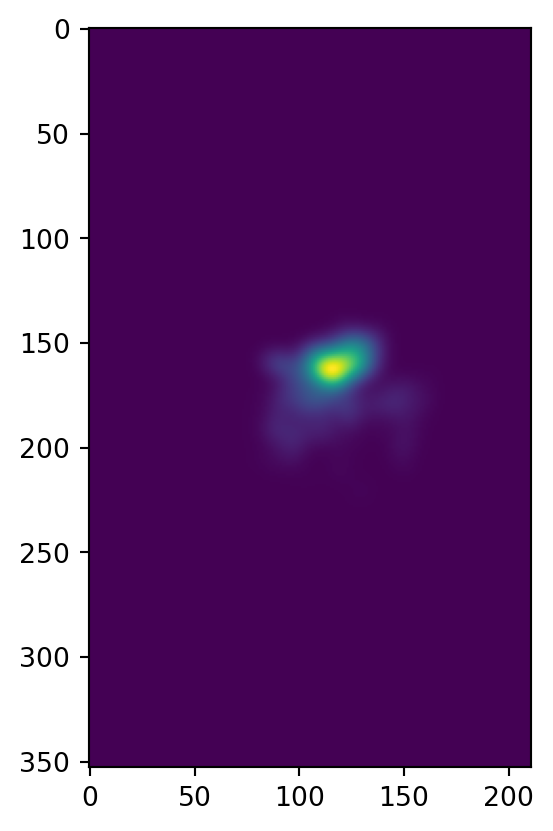
First, we’ll bin things into a 2D grid:
sigma = 500 # meters
delta = 100 # meters
xy = np.array([x.coords[0] for x in gdf['start_points']])
extent = [np.min(xy[:,1]), np.max(xy[:,1]), np.min(xy[:,0]), np.max(xy[:,0])]
xx = np.linspace(extent[2], extent[3], int(abs(extent[3]-extent[2])/delta))
yy = np.linspace(extent[0], extent[1], int(abs(extent[1]-extent[0])/delta))
heatmap, xedges, yedges = np.histogram2d(xy[:,0], xy[:,1], bins=[xx, yy])
plt.imshow(heatmap)
Then, we smooth it: (question: how’s this relate to how we did KDE above?)
Better plotting
There’s various ways to do this. One is to use contextily as demonstrated here. First, we need to convert to “EPSG:3857 (Web Mercator)”:
xpts = gpd.points_from_xy(xx, np.full((len(xx),), np.mean(yy)), crs="EPSG:5070").to_crs("EPSG:3857")
ypts = gpd.points_from_xy(np.full((len(yy),), np.mean(xx)), yy, crs="EPSG:5070").to_crs("EPSG:3857")
gpd.GeoDataFrame(geometry=gpd.points_from_xy(
np.concat([xx, np.full((len(yy),), np.mean(xx))]),
np.concat([np.full((len(xx),), np.mean(yy)), yy])),
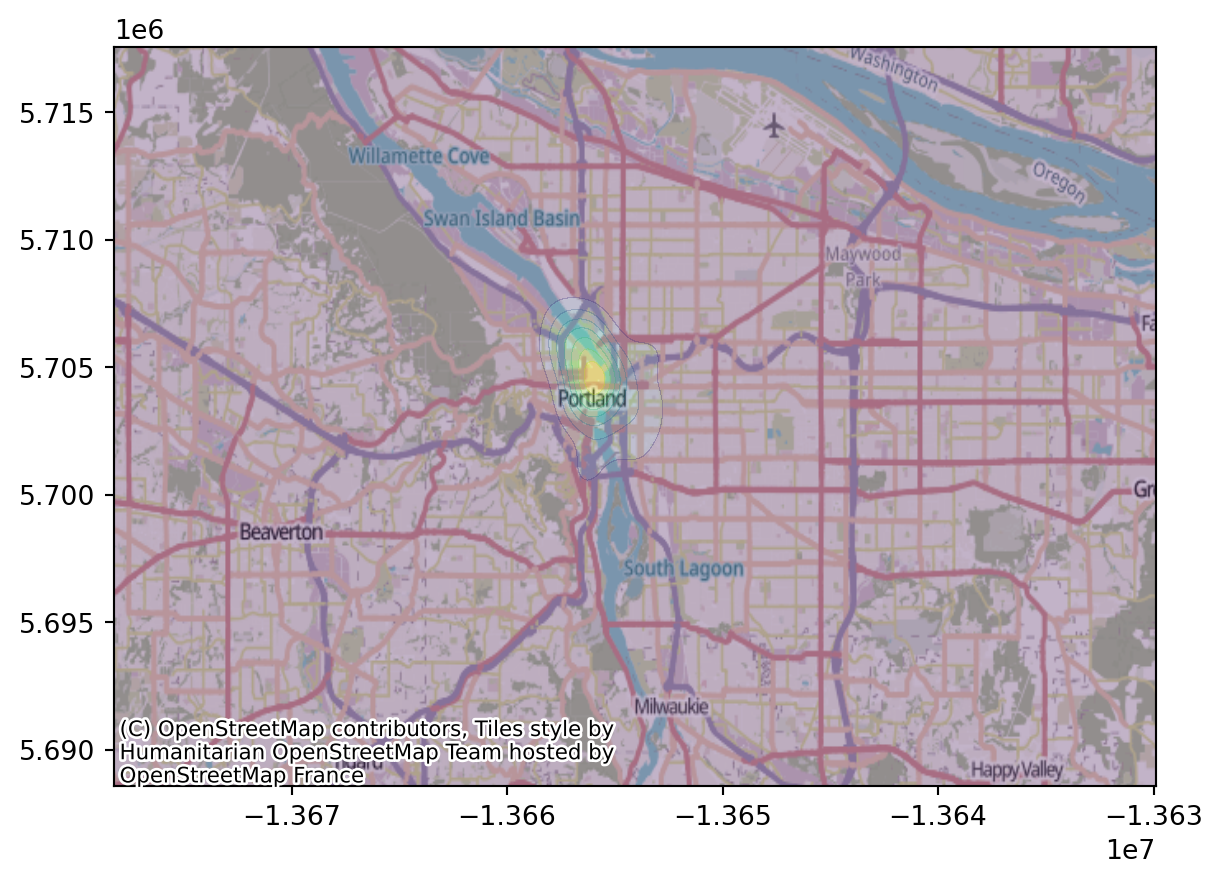
crs="EPSG:5070").to_crs("EPSG:3857").explore()Now, we can superimpose on the OSM map:
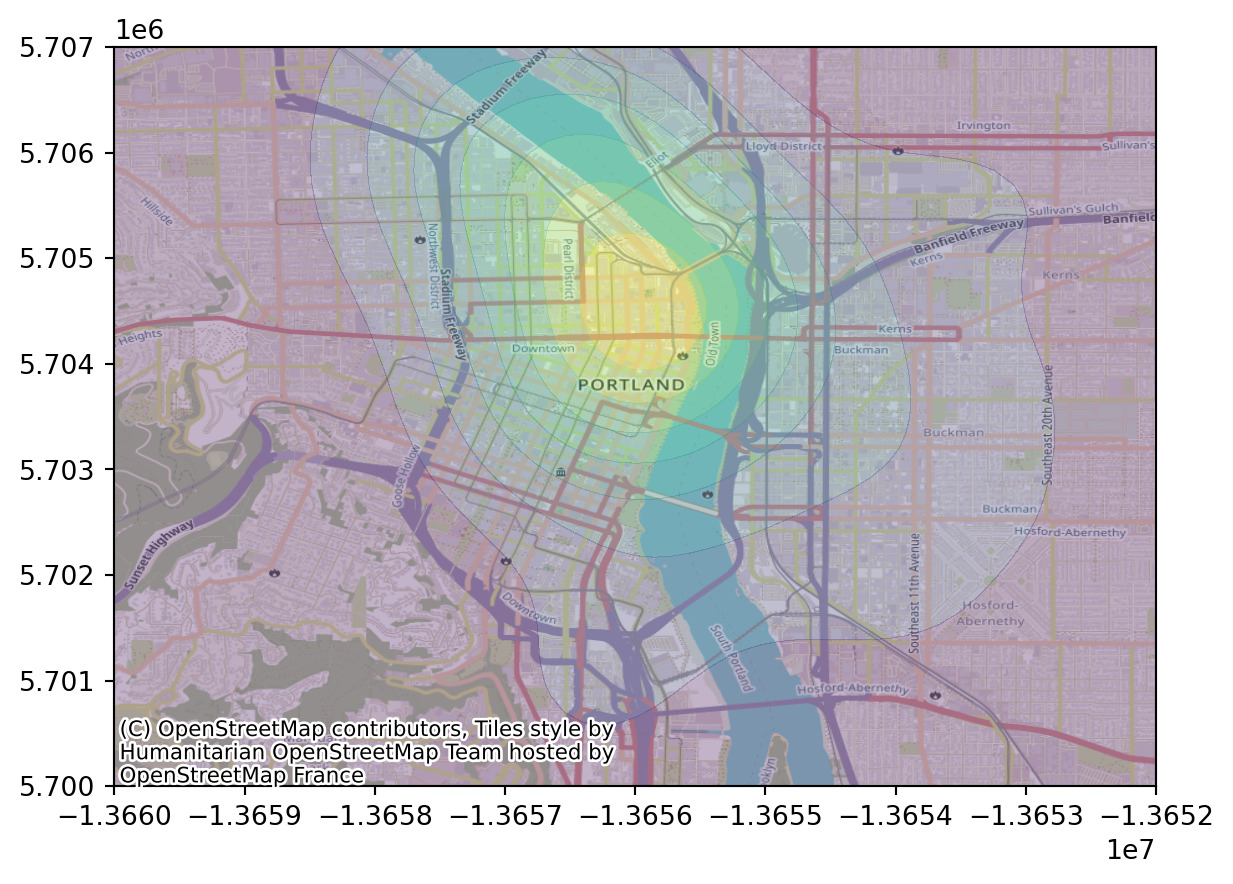
Let’s zoom in:
Bandwidth is important
Question:
Above I set:
sigma = 500 # meters
delta = 100 # metersWhat are these numbers, in words, and are these good choices?
Exercise:
Explore different choices of these two parameters. To do this:
- Write a function that takes in
sigmaanddeltaand makes a plot. - Play around with it.
Choosing a bandwidth
How to pick a bandwidth?
There are various automatic methods.
But… crossvalidation!
For each bandwidth:
- Fit using most of the data,
- and predict the remaining data.
- Do this a bunch and return an estimate of goodness-of-fit.
… but wait, what’s this mean, here?
Revised:
For each bandwidth:
- Fit the smooth using most of the data,
- predict the density at the locations of the remaining data,
- and return the mean log density at “true” points.
Why log?:
If \(f\) and \(g\) are probability distributions, and \(x_1, \ldots, x_n\) are drawn from \(f\), then \[ L(g) = \sum_{i=1}^n \log g(x_i) \lessapprox L(f) , \] i.e., is maximized for \(g=f\). (see: cross entropy)