Case study: Groundwater monitoring
2026-02-16
Wastewater in Eugene
Our wastewater (sewer plus stormwater) in Eugene and Springfield is treated by the Metropolitan Wastewater Management Commission, in part at the Biosolids Management Facility (see slideshow on that page).
As part of their operating requirements, they monitor groundwater for potential contamination: in 26 monitoring wells, since 1988.
Their biggest question: how to best detect if the facility is affecting the groundwater?
Note: the MWMC is very much on top of this, and has no evidence of groundwater contamination. This is being presented for pedagogical purposes.
What is measured?
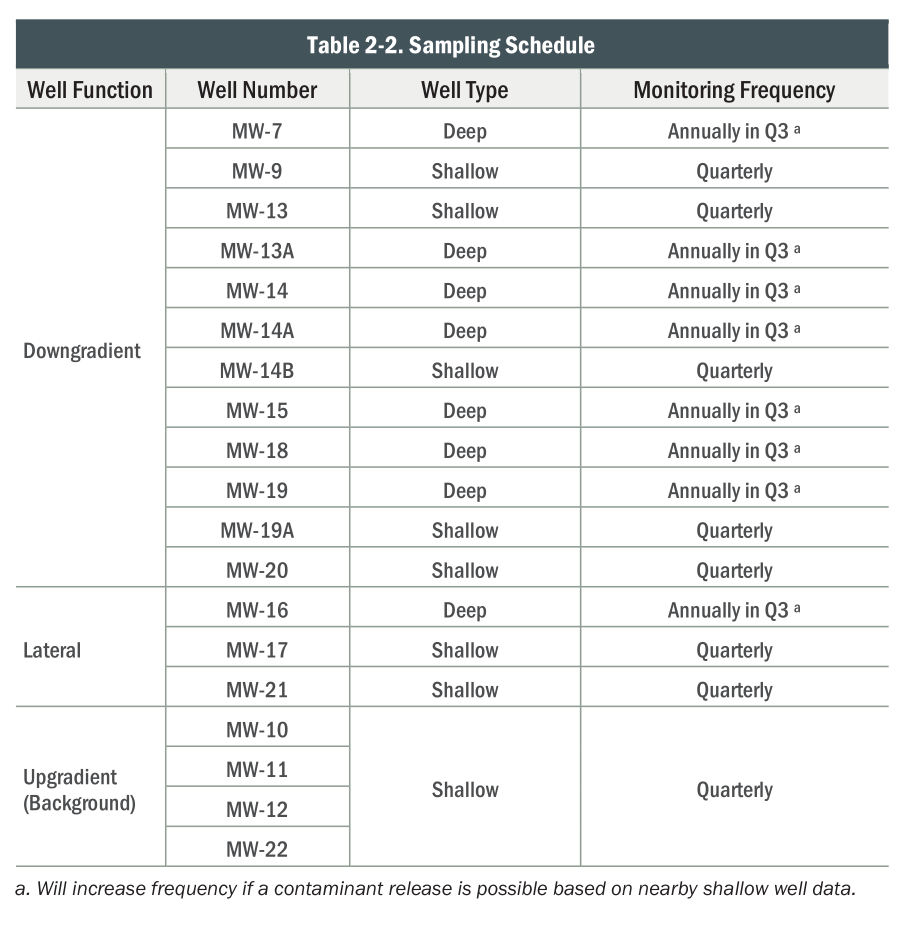
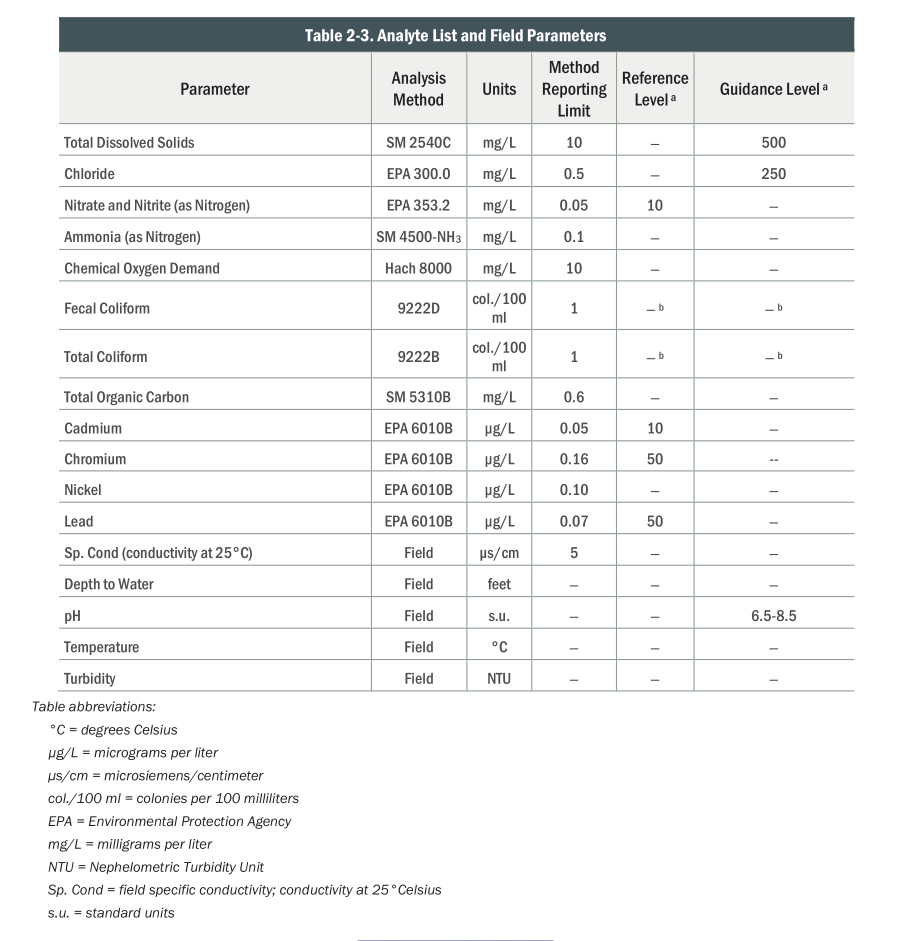 )
)
Overview of the data:
We’ve got:
Information about the wells: BMF_Well_details_2023_GWMP.csv
The actual measurements: Eugene_BiosolidsManagementFacility_GroundwaterData.csv
Set-up
“Look at the data”, part 1
Game plan:
- How many wells are there?
- How many observations do we have from each, of what quantities, and over what time periods?
- Where are the wells, and what do we know about them?
Preliminary answers above; but let’s check this!
Measurements
- Measurements often run into detection limits; this is conveyed as “censoring status”.
well.nameis the name of the location with the first year it was in operation appended
$ head data/Eugene_BiosolidsManagementFacility_GroundwaterData.csv | tr ',' '\t'
"well.name" "SampledDate" "SampleNumber" "ParameterName" "ParameterUnits" "result" "Method.Detection.Limit" "Reporting.limit" "censoring.status" "Easting" "Northing"
"MW-13-1988" 1988-01-25 "880050" "Ammonia" "mg/L" 0.1 NA NA "left.censored" 4218262.9 910814.599999998
"MW-13A-1988" 1988-01-25 "880052" "Ammonia" "mg/L" 0.1 NA NA "left.censored" 4218304.4 910813.2
"MW-14-1988" 1988-01-25 "880054" "Ammonia" "mg/L" 0.1 NA NA "left.censored" 4217612.3 910814.700000002
"MW-14A-1988" 1988-01-25 "880055" "Ammonia" "mg/L" 0.1 NA NA "left.censored" 4217622.5 910814.499999998
"MW-14B-1988" 1988-01-25 "880058" "Ammonia" "mg/L" 0.1 NA NA "left.censored" 4217612.9 910804.299999995
"MW-13-1988" 1988-01-25 "880050" "Cadmium" "µg/L" 1 NA NA "left.censored" 4218262.9 910814.599999998
"MW-13A-1988" 1988-01-25 "880052" "Cadmium" "µg/L" 1 NA NA "left.censored" 4218304.4 910813.2
"MW-14-1988" 1988-01-25 "880054" "Cadmium" "µg/L" 1 NA NA "left.censored" 4217612.3 910814.700000002
"MW-14A-1988" 1988-01-25 "880055" "Cadmium" "µg/L" 1 NA NA "left.censored" 4217622.5 910814.499999998Pull in the data
| well_name | SampledDate | SampleNumber | ParameterName | ParameterUnits | result | Method_Detection_Limit | Reporting_limit | censoring_status | Easting | Northing | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | MW-13-1988 | 1988-01-25 | 880050 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4218262.9 | 910814.6 |
| 1 | MW-13A-1988 | 1988-01-25 | 880052 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4218304.4 | 910813.2 |
| 2 | MW-14-1988 | 1988-01-25 | 880054 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4217612.3 | 910814.7 |
| 3 | MW-14A-1988 | 1988-01-25 | 880055 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4217622.5 | 910814.5 |
| 4 | MW-14B-1988 | 1988-01-25 | 880058 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4217612.9 | 910804.3 |
| ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... | ... |
| 49552 | MW-9-1996 | 2026-01-21 | 012126-01-03 | WaterTableElevation | feet above mean sea level | 361.44 | <NA> | <NA> | not.censored | 4217256.3 | 909769.9 |
| 49553 | MW-19A-1988 | 2026-01-21 | 012126-01-04 | WaterTableElevation | feet above mean sea level | 361.72 | <NA> | <NA> | not.censored | 4217612.3 | 910354.1 |
| 49554 | MW-13-2024 | 2026-01-21 | 012126-01-05 | WaterTableElevation | feet above mean sea level | <NA> | <NA> | <NA> | not.censored | 4218339.234158 | 910812.64945 |
| 49555 | MW-14B-1988 | 2026-01-21 | 012126-01-06 | WaterTableElevation | feet above mean sea level | 360.56 | <NA> | <NA> | not.censored | 4217612.9 | 910804.3 |
| 49556 | MW-20-1988 | 2026-01-21 | 012126-01-07 | WaterTableElevation | feet above mean sea level | 361.5 | <NA> | <NA> | not.censored | 4217582.6 | 909534.0 |
49557 rows × 11 columns
What’s measured?
| count | mean | std | min | 25% | 50% | 75% | max | |
|---|---|---|---|---|---|---|---|---|
| ParameterName | ||||||||
| Ammonia | 3308.0 | 0.103789 | 0.03004 | 0.021 | 0.1 | 0.1 | 0.1 | 0.6 |
| Cadmium | 2508.0 | 0.390975 | 0.459542 | 0.0027 | 0.0075 | 0.2 | 1.0 | 5.0 |
| ChemicalOxygenDemand | 2489.0 | 8.962302 | 3.715514 | 3.6 | 5.0 | 10.0 | 10.0 | 96.0 |
| Chloride | 3316.0 | 5.562295 | 3.830561 | 0.5 | 3.8 | 5.0 | 6.9 | 49.0 |
| Chromium | 2510.0 | 2.394473 | 2.252434 | 0.045 | 0.511 | 2.0 | 5.0 | 56.0 |
| Conductivity | 3522.0 | 344.5477 | 143.069563 | 92.0 | 248.0 | 308.5 | 410.0 | 950.0 |
| Copper | 2520.0 | 2.897059 | 2.50814 | 0.039 | 0.3205 | 5.0 | 5.0 | 46.0 |
| FecalColiform | 3356.0 | 3.158522 | 25.10456 | 1.0 | 1.0 | 1.0 | 1.0 | 800.0 |
| Lead | 2502.0 | 2.105006 | 2.237623 | 0.002 | 0.0294 | 2.0 | 5.0 | 16.0 |
| Nickel | 2507.0 | 4.459762 | 4.080386 | 0.0087 | 0.456 | 5.0 | 10.0 | 13.0 |
| Nitrate | 3350.0 | 3.689657 | 2.205737 | 0.02 | 2.456 | 3.4 | 4.6 | 15.0 |
| Temperature | 223.0 | 13.675336 | 1.824945 | 8.6 | 12.6 | 13.5 | 14.8 | 18.8 |
| TotalColiform | 3346.0 | 6.614764 | 61.624235 | 1.0 | 1.0 | 1.0 | 1.0 | 2100.0 |
| TotalDissolvedSolids | 3358.0 | 231.526504 | 80.129682 | 60.0 | 180.0 | 214.0 | 270.0 | 880.0 |
| TotalOrganicCarbon | 2442.0 | 1.319935 | 1.039013 | 0.2 | 0.5 | 0.9 | 2.0 | 22.0 |
| WaterTableElevation | 3539.0 | 358.891123 | 3.812151 | 351.66 | 355.35 | 359.42 | 361.865 | 370.09 |
| Zinc | 2504.0 | 5.654278 | 4.790525 | 0.07 | 0.5735 | 10.0 | 10.0 | 20.0 |
| pH | 2250.0 | 6.945916 | 0.243518 | 5.94 | 6.81 | 7.0 | 7.1 | 8.4 |
How many things are measured?
Tabular form:
| ParameterName | Ammonia | Cadmium | ChemicalOxygenDemand | Chloride | Chromium | Conductivity | Copper | FecalColiform | Lead | Nickel | Nitrate | Temperature | TotalColiform | TotalDissolvedSolids | TotalOrganicCarbon | WaterTableElevation | Zinc | pH |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| well_name | ||||||||||||||||||
| MW-10-1983 | 44 | 47 | 45 | 47 | 47 | 47 | 47 | 47 | 47 | 47 | 43 | 0 | 44 | 45 | 43 | 48 | 47 | 0 |
| MW-10-1996 | 132 | 87 | 86 | 131 | 86 | 138 | 87 | 132 | 86 | 86 | 131 | 17 | 131 | 131 | 86 | 139 | 86 | 118 |
| MW-11-1983 | 47 | 48 | 48 | 47 | 48 | 49 | 48 | 49 | 48 | 48 | 48 | 0 | 46 | 49 | 45 | 49 | 48 | 0 |
| MW-11-1996 | 130 | 85 | 86 | 130 | 85 | 140 | 84 | 130 | 84 | 84 | 131 | 14 | 130 | 132 | 84 | 140 | 83 | 117 |
| MW-12-1983 | 46 | 47 | 47 | 48 | 47 | 46 | 47 | 48 | 47 | 47 | 48 | 0 | 47 | 48 | 44 | 48 | 47 | 0 |
| MW-12-1996 | 125 | 86 | 83 | 126 | 86 | 140 | 85 | 124 | 84 | 85 | 126 | 14 | 125 | 126 | 83 | 142 | 85 | 122 |
| MW-13-1988 | 141 | 97 | 98 | 140 | 97 | 149 | 97 | 147 | 97 | 97 | 141 | 1 | 144 | 142 | 93 | 152 | 97 | 80 |
| MW-13-2018 | 26 | 27 | 26 | 26 | 28 | 32 | 30 | 27 | 28 | 29 | 27 | 2 | 27 | 26 | 27 | 31 | 29 | 32 |
| MW-13-2024 | 7 | 6 | 6 | 6 | 6 | 7 | 6 | 7 | 6 | 6 | 6 | 7 | 7 | 6 | 6 | 7 | 6 | 7 |
| MW-13A-1988 | 170 | 129 | 128 | 171 | 130 | 178 | 129 | 173 | 129 | 129 | 174 | 7 | 172 | 173 | 125 | 177 | 129 | 110 |
| MW-14-1988 | 169 | 125 | 125 | 170 | 125 | 177 | 126 | 172 | 125 | 125 | 173 | 8 | 172 | 174 | 122 | 179 | 125 | 110 |
| MW-14A-1988 | 179 | 132 | 132 | 179 | 132 | 184 | 133 | 181 | 132 | 132 | 182 | 6 | 181 | 181 | 128 | 187 | 132 | 116 |
| MW-14B-1988 | 180 | 138 | 136 | 182 | 139 | 193 | 141 | 184 | 138 | 140 | 182 | 16 | 184 | 183 | 136 | 192 | 140 | 124 |
| MW-15-1988 | 163 | 124 | 124 | 164 | 123 | 174 | 124 | 166 | 123 | 123 | 165 | 6 | 166 | 166 | 122 | 177 | 124 | 110 |
| MW-16-1988 | 169 | 129 | 130 | 170 | 131 | 178 | 133 | 171 | 129 | 130 | 172 | 8 | 171 | 174 | 126 | 181 | 130 | 112 |
| MW-17-1988 | 182 | 138 | 137 | 180 | 136 | 192 | 136 | 184 | 135 | 137 | 181 | 14 | 184 | 181 | 134 | 193 | 137 | 126 |
| MW-18-1988 | 181 | 136 | 136 | 182 | 136 | 188 | 137 | 183 | 136 | 136 | 185 | 6 | 183 | 183 | 133 | 189 | 135 | 118 |
| MW-19-1988 | 165 | 125 | 124 | 167 | 125 | 175 | 125 | 167 | 125 | 125 | 170 | 6 | 167 | 169 | 123 | 174 | 125 | 108 |
| MW-19A-1988 | 175 | 135 | 131 | 176 | 134 | 197 | 135 | 180 | 134 | 133 | 179 | 19 | 179 | 176 | 130 | 201 | 134 | 136 |
| MW-20-1988 | 183 | 136 | 134 | 179 | 138 | 199 | 139 | 189 | 139 | 136 | 186 | 18 | 189 | 182 | 135 | 202 | 135 | 137 |
| MW-21-1988 | 178 | 136 | 135 | 179 | 136 | 187 | 137 | 180 | 136 | 136 | 179 | 15 | 180 | 181 | 133 | 185 | 136 | 121 |
| MW-22-1988 | 172 | 133 | 130 | 172 | 133 | 184 | 133 | 175 | 133 | 135 | 173 | 19 | 176 | 175 | 129 | 186 | 133 | 117 |
| MW-7-1983 | 45 | 48 | 48 | 46 | 48 | 49 | 48 | 40 | 48 | 48 | 46 | 0 | 45 | 48 | 45 | 49 | 48 | 0 |
| MW-7-1996 | 123 | 78 | 78 | 123 | 78 | 126 | 78 | 123 | 78 | 78 | 123 | 5 | 123 | 126 | 78 | 128 | 78 | 106 |
| MW-9-1983 | 47 | 48 | 48 | 47 | 48 | 48 | 48 | 48 | 48 | 48 | 47 | 0 | 44 | 48 | 45 | 49 | 48 | 0 |
| MW-9-1996 | 129 | 88 | 88 | 128 | 88 | 145 | 87 | 129 | 87 | 87 | 132 | 15 | 129 | 133 | 87 | 141 | 87 | 123 |
When was each well used?
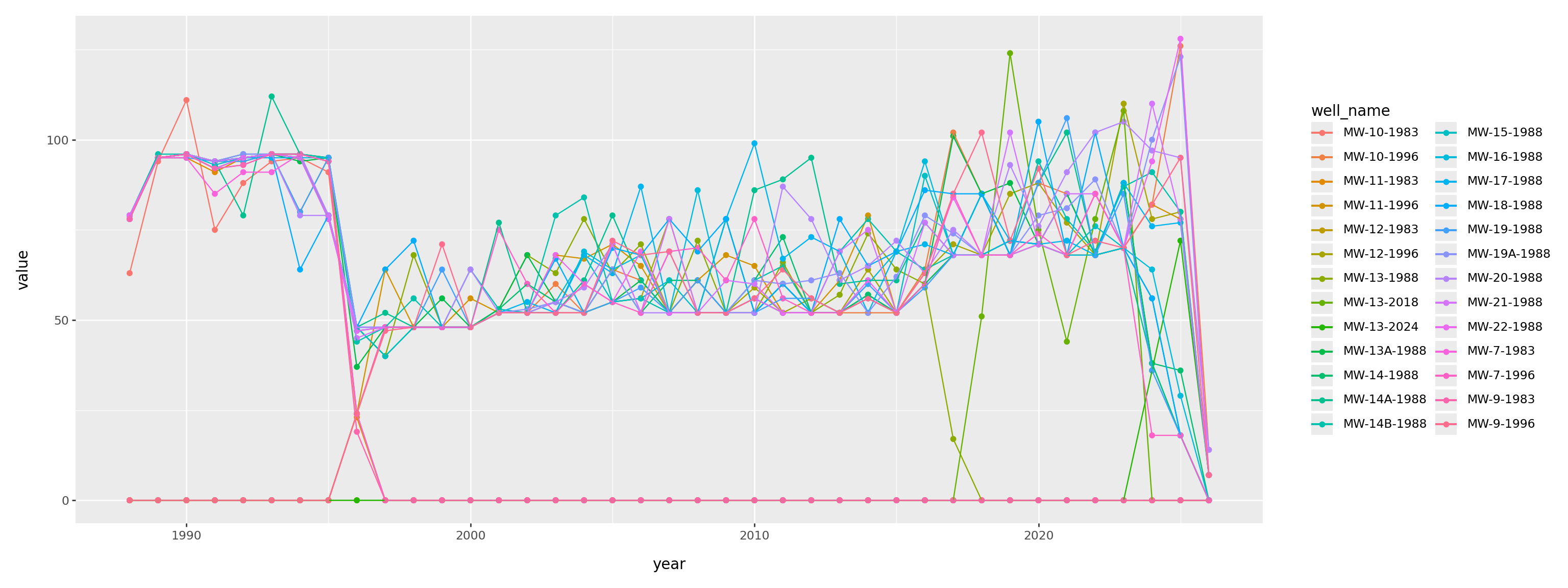
When was each thing measured?
(
pd.crosstab(gw['ParameterName'], gw['SampledDate'].dt.year)
.reset_index()
.melt(id_vars=['ParameterName'])
.assign(year=lambda df: pd.to_numeric(df['SampledDate']))
>>
p9.ggplot(p9.aes(x='year', y='value', color='ParameterName'))
+ p9.geom_point() + p9.geom_line()
+ p9.labs(y='number of measurements')
+ p9.theme(figure_size=(16,6))
)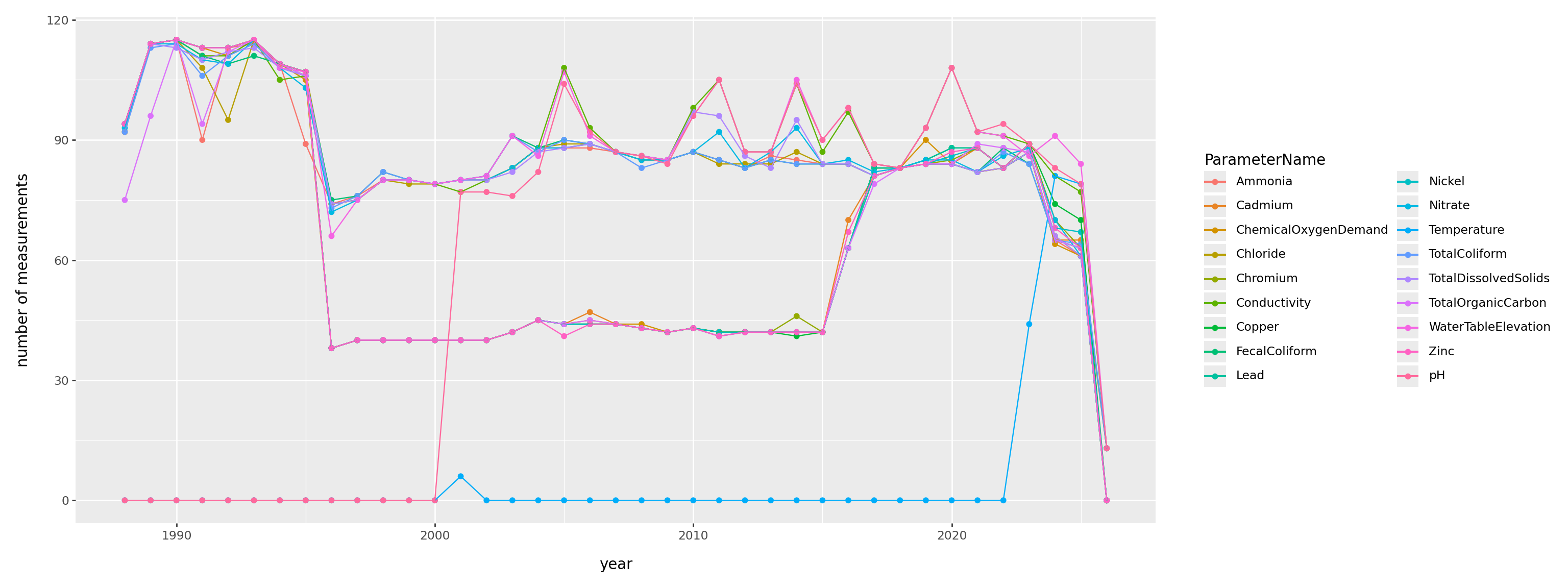
Summary:
How many observations do we have from each, of what quantities, and over what time periods?
Exercise.
Where are the wells?
Well info
$ cat data/BMF_Well_details_2023_GWMP.csv | tr ',' '\t'
LocationName well.orientation well.depth.category screened.interval.feet
MW-7 downgradient deep 42 to 52
MW-9 downgradient shallow 15 to 25
MW-13A downgradient deep 30 to 50
MW-14 downgradient deep 55 to 75
MW-14A downgradient deep 30 to 50
MW-14B downgradient shallow 10 to 25
MW-15 downgradient deep 30 to 50
MW-18 downgradient deep 30 to 50
MW-19 downgradient deep 30 to 50
MW-19A downgradient shallow 10 to 25
MW-20 downgradient shallow 10 to 25
MW-16 lateral deep 30 to 50
MW-17 lateral shallow 10 to 25
MW-21 lateral shallow 10 to 25
MW-10 upgradient shallow 15 to 25
MW-11 upgradient shallow 15 to 25
MW-12 upgradient shallow 15 to 25
MW-22 upgradient shallow 10 to 25
MW-13-2024 downgradient shallow 10 to 25Get per-well summaries from this
We’d like the location to be in the per-well table, so let’s pull this out of the “measurements” file first:
gw_wells = (
gw.groupby(['well_name', 'Easting', 'Northing'])
.aggregate(
n = ('SampleNumber', lambda x: len(x)),
)
.reset_index()
.assign(
LocationName = lambda df: "MW-" + df['well_name'].str.extract("-([0-9A-Z]*)-"),
FirstYear = lambda df: df['well_name'].str.extract("-([0-9A-Z]*)$")
)
)
gw_wells.loc[gw_wells['well_name'] == "MW-13-2024", 'LocationName'] = "MW-13-2024"Well information
| LocationName | well_orientation | well_depth_category | screened_interval_feet | well_name | Easting | Northing | n | FirstYear | |
|---|---|---|---|---|---|---|---|---|---|
| 0 | MW-10 | upgradient | shallow | 15 to 25 | MW-10-1983 | 4219307.315854 | 909474.081725 | 735 | 1983 |
| 1 | MW-10 | upgradient | shallow | 15 to 25 | MW-10-1996 | 4217879.6 | 908303.2 | 1890 | 1996 |
| 2 | MW-11 | upgradient | shallow | 15 to 25 | MW-11-1983 | 4217171.326475 | 908754.90989 | 765 | 1983 |
| 3 | MW-11 | upgradient | shallow | 15 to 25 | MW-11-1996 | 4217227.5 | 908744.7 | 1869 | 1996 |
| 4 | MW-12 | upgradient | shallow | 15 to 25 | MW-12-1983 | 4219118.324783 | 908681.318636 | 752 | 1983 |
| 5 | MW-12 | upgradient | shallow | 15 to 25 | MW-12-1996 | 4219546.5 | 908262.3 | 1847 | 1996 |
| 6 | MW-13 | <NA> | <NA> | <NA> | MW-13-1988 | 4218262.9 | 910814.6 | 2010 | 1988 |
| 7 | MW-13 | <NA> | <NA> | <NA> | MW-13-2018 | 4218262.9 | 910814.6 | 480 | 2018 |
| 8 | MW-13-2024 | downgradient | shallow | 10 to 25 | MW-13-2024 | 4218339.234158 | 910812.64945 | 115 | 2024 |
| 9 | MW-13A | downgradient | deep | 30 to 50 | MW-13A-1988 | 4218304.4 | 910813.2 | 2533 | 1988 |
| 10 | MW-14 | downgradient | deep | 55 to 75 | MW-14-1988 | 4217612.3 | 910814.7 | 2502 | 1988 |
| 11 | MW-14A | downgradient | deep | 30 to 50 | MW-14A-1988 | 4217622.5 | 910814.5 | 2629 | 1988 |
| 12 | MW-14B | downgradient | shallow | 10 to 25 | MW-14B-1988 | 4217612.9 | 910804.3 | 2728 | 1988 |
| 13 | MW-15 | downgradient | deep | 30 to 50 | MW-15-1988 | 4217592.3 | 909964.4 | 2444 | 1988 |
| 14 | MW-16 | lateral | deep | 30 to 50 | MW-16-1988 | 4217592.7 | 909113.7 | 2544 | 1988 |
| 15 | MW-17 | lateral | shallow | 10 to 25 | MW-17-1988 | 4219020.1 | 910333.0 | 2707 | 1988 |
| 16 | MW-18 | downgradient | deep | 30 to 50 | MW-18-1988 | 4217912.6 | 910784.0 | 2683 | 1988 |
| 17 | MW-19 | downgradient | deep | 30 to 50 | MW-19-1988 | 4217612.7 | 910364.2 | 2465 | 1988 |
| 18 | MW-19A | downgradient | shallow | 10 to 25 | MW-19A-1988 | 4217612.3 | 910354.1 | 2684 | 1988 |
| 19 | MW-20 | downgradient | shallow | 10 to 25 | MW-20-1988 | 4217582.6 | 909534.0 | 2756 | 1988 |
| 20 | MW-21 | lateral | shallow | 10 to 25 | MW-21-1988 | 4218063.0 | 908984.0 | 2670 | 1988 |
| 21 | MW-22 | upgradient | shallow | 10 to 25 | MW-22-1988 | 4218563.1 | 908764.3 | 2608 | 1988 |
| 22 | MW-7 | downgradient | deep | 42 to 52 | MW-7-1983 | 4217236.222717 | 910903.762698 | 749 | 1983 |
| 23 | MW-7 | downgradient | deep | 42 to 52 | MW-7-1996 | 4217291.5 | 910895.9 | 1730 | 1996 |
| 24 | MW-9 | downgradient | shallow | 15 to 25 | MW-9-1983 | 4217197.302791 | 909779.024971 | 759 | 1983 |
| 25 | MW-9 | downgradient | shallow | 15 to 25 | MW-9-1996 | 4217256.3 | 909769.9 | 1903 | 1996 |
Mapping
Let’s use folium. Looking at the documentation, we need a location in “Latitude and Longitude of Map (Northing, Easting)”.
Easting and Northing coordinates are provided in the ESPG: 2270 Coordinate Reference System (NAD83 / Oregon South (ft))
So first: find the center of the well locations, and project it to lat/long.
def transform(easting, northing):
crs = pyproj.CRS.from_epsg(2270)
latlon = pyproj.CRS(proj='latlon', datum='WGS84')
transformer = pyproj.Transformer.from_crs(crs, latlon)
y, x = transformer.transform(easting, northing)
return (x, y)
center = transform(wells['Easting'].mean(), wells['Northing'].mean())
center(44.132343700797044, -123.17902808886248)The wells:
Let’s get some more info on that:
After some iteration:
import matplotlib
def add_marker(m, row=None, tooltip=None, location=None, **kwargs):
defaults = {
'radius': 10,
'fill_color': "orange",
'fill_opacity': 0.4,
'color': 'black',
'weight': 1
}
defaults.update(kwargs)
if location is None:
location = transform(row.Easting, row.Northing)
if tooltip is None and row is not None:
tooltip = row.LocationName
return folium.CircleMarker(
location=location,
tooltip=tooltip,
**defaults
).add_to(m)
def make_colormap(x, xmin=None, xmax=None, mapname='viridis'):
cmap = matplotlib.colormaps[mapname]
if xmin is None:
xmin = np.min(x)
if xmax is None:
xmax = np.max(x)
assert xmax > xmin, f"Must have xmin < xmax (got {xmin}, {xmax})."
def f(x):
colors = cmap((x - xmin)/(xmax - xmin))
if isinstance(colors, tuple):
colors = [colors]
return [matplotlib.colors.to_hex(u) for u in colors]
return f
well_bbox = (
transform(wells['Easting'].min(), wells['Northing'].min()),
transform(wells['Easting'].min(), wells['Northing'].max()),
transform(wells['Easting'].max(), wells['Northing'].max()),
transform(wells['Easting'].max(), wells['Northing'].min()),
)
def add_legend(m, colors, k=8, title=None):
dy = (well_bbox[1][0] - well_bbox[0][0])/k
pos = list(well_bbox[2])
def text(pos, label):
folium.Marker(
pos,
icon=folium.DivIcon(
icon_size=(100,30),
icon_anchor=(-15,15),
html=f'<div style="font-size: 15pt">{label}</div>',
)
).add_to(m)
if title is not None:
text(pos, title)
pos[0] -= dy
for n in colors:
add_marker(m, location=pos, fill_color=colors[n]).add_to(m)
label = n if type(n) is str else 'NA'
text(pos, label)
pos[0] -= dyWell orientation:
orientation_colors = {
'upgradient' : "#66c2a5",
'downgradient' : "#8da0cb",
'lateral' : "#fc8d62",
pd.NA : "#666666",
}
m = folium.Map(location=center, zoom_start=16)
add_legend(m, orientation_colors)
for r in wells.itertuples():
add_marker(m, r,
fill_color=orientation_colors[r.well_orientation],
)
mWell depth:
Summary
Where are the wells, and what do we know about them?
Exercise
“Look at the data”, part 2
Game plan:
Do the measurements differ most:
- between wells?
- between types of well? (up/downgradient, shallow/deep)
- between years?
- between seasons?
- all/some of the above?
Do nearby wells get similar measurements?
All the data, from 20,000 feet
/home/peter/micromamba/envs/ds435/lib/python3.13/site-packages/plotnine/layer.py:374: PlotnineWarning: geom_point : Removed 7 rows containing missing values.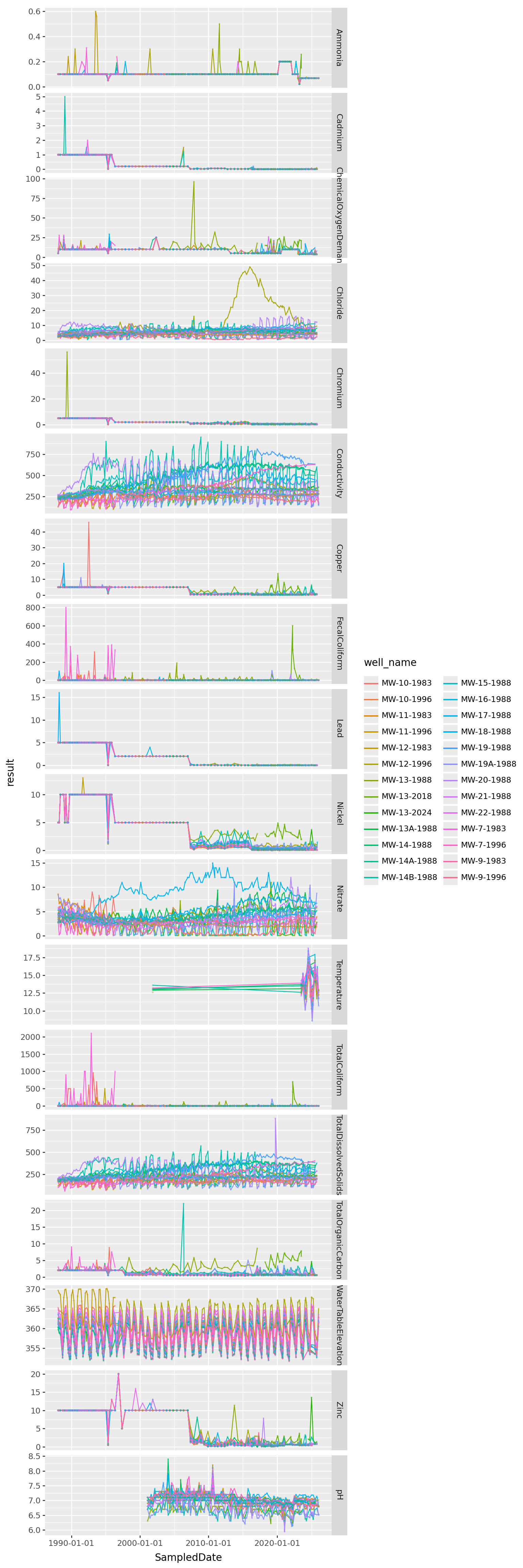
Observations?
What features do we see here and what do those features suggest doing?
Exercise:
Filter out the obvious outliers (eg Chromium, can’t tell what’s going on).
Which variables are most important to the audience of the report?
Consider removing wells or variables for which we don’t have complete observations: for instance, Temperature.
Conductivity:
- does the location relative to the plant,
- or type of well
- impact whether it’s increasing/decreasing long term,
- or whether it’s oscilating.
Are the oscillations in various variables always going along with water level/season?
Chloride: what happened with that one well that had very large values for a while?
Set-up:
Let’s add well categories to the measurement data:
| well_name | SampledDate | SampleNumber | ParameterName | ParameterUnits | result | Method_Detection_Limit | Reporting_limit | censoring_status | Easting | Northing | well_orientation | well_depth_category | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | MW-13-1988 | 1988-01-25 | 880050 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4218262.9 | 910814.6 | <NA> | <NA> |
| 1 | MW-13A-1988 | 1988-01-25 | 880052 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4218304.4 | 910813.2 | downgradient | deep |
| 2 | MW-14-1988 | 1988-01-25 | 880054 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4217612.3 | 910814.7 | downgradient | deep |
| 3 | MW-14A-1988 | 1988-01-25 | 880055 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4217622.5 | 910814.5 | downgradient | deep |
| 4 | MW-14B-1988 | 1988-01-25 | 880058 | Ammonia | mg/L | 0.1 | <NA> | <NA> | left.censored | 4217612.9 | 910804.3 | downgradient | shallow |
Outliers?
Here’s Chromium:
Ways to deal with outliers in visualization
- Change the axis limits to zoom in on what you want.
- Transform the variable (e.g., with
log). - Programatically decide which ones are outliers and remove them.
- Either the second or third methods sets you up for downstream analysis.
- A transformation is better if it gives you “the right scale to look at the data on”.
- The third method is good if you think the “outliers” are produced by some other process that is not of interest (for instance, measurement error).
- A good generic way to do (3) is by computing \(z\) scores:
z = (value - center)/scale.
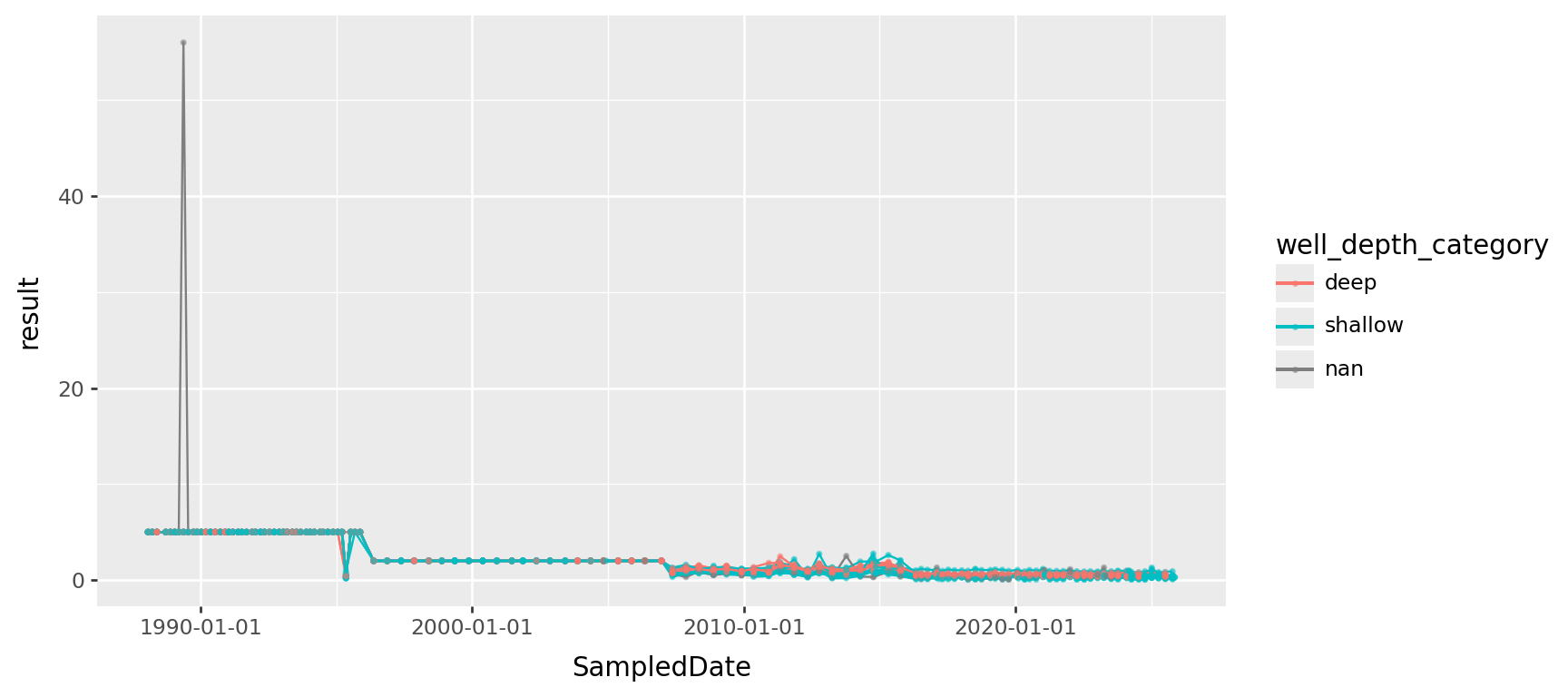
Chromium, transformed
(
gw
.query("ParameterName == 'Chromium'")
>> p9.ggplot(p9.aes(x="SampledDate", y="result", color="well_depth_category", group="well_name"))
+ p9.geom_line() + p9.geom_point(size=0.5, alpha=0.5)
+ p9.labs(title="Chromium", y="concentration (µg/L)")
+ p9.scale_y_log10()
) + p9.theme(figure_size=(9,4))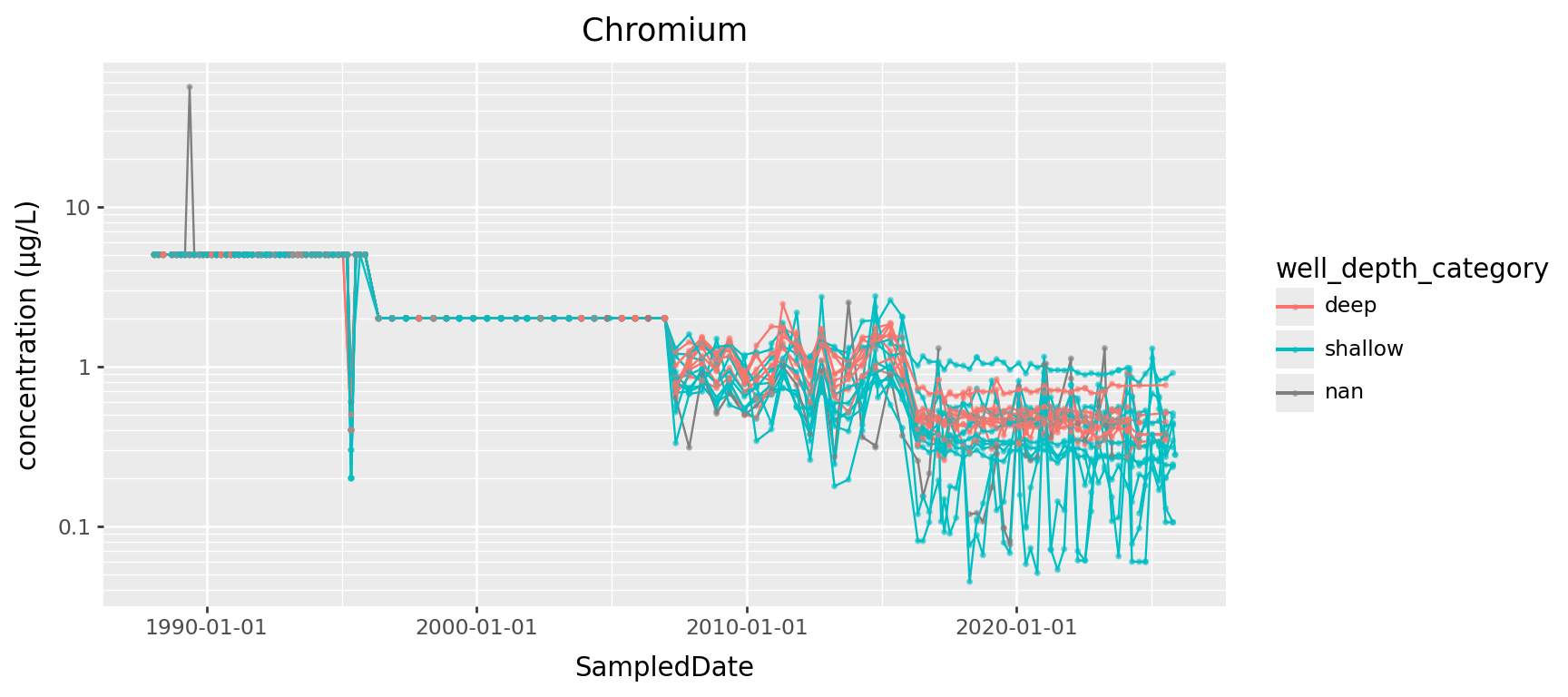
Chromium, “identifying outliers”?
z = (value - center) / scale
Question: what to use for “center” and “scale”? Median+IQR or mean+SD? Within groups? Which groups?
- Definitely group by year in this case, since things change.
- Usually I’d prefer “median and IQR”, but for many years the IQR is zero.
chromium = (
gw.query("ParameterName == 'Chromium'")
.assign(
mean = lambda df: df['result'].groupby(df['SampledDate'].dt.year).transform('mean'),
std = lambda df: df['result'].groupby(df['SampledDate'].dt.year).transform('std'),
z = lambda df: (df['result'] - df['mean']) / ( df['std'] + (df['std'] == 0) ),
)
)
chromium >> p9.ggplot(p9.aes(x='z')) + p9.geom_histogram(bins=40)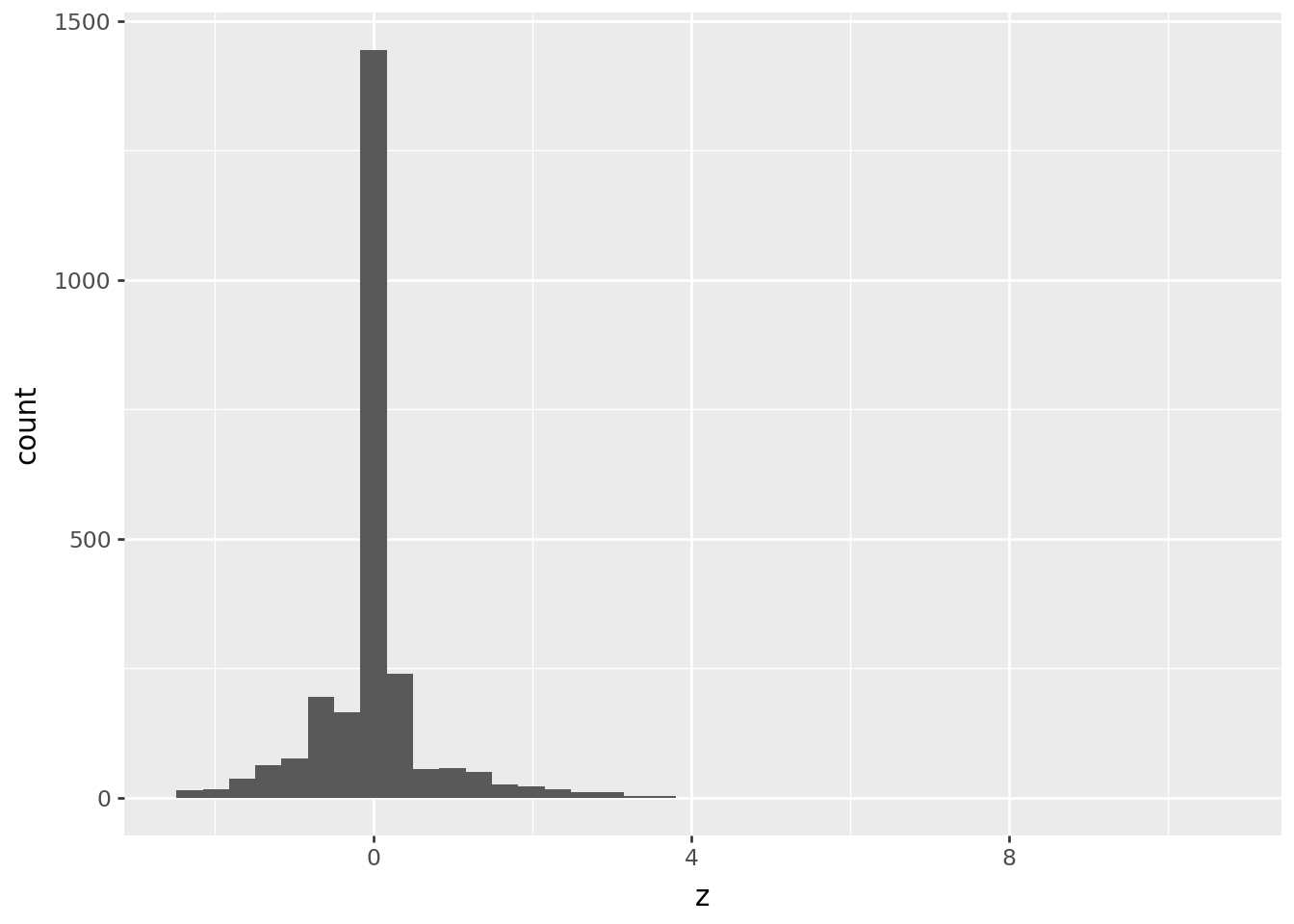
How’s that look?
The cutoff for “outlier” in terms of \(z\) scores should always be made looking at the data, but often between 3 and 5 is good, if this is a reasonable approach.
p1 = (
chromium
>>
p9.ggplot(p9.aes(x="SampledDate", y="result", group="well_name"))
+ p9.geom_line(alpha=0.5)
+ p9.geom_point(mapping=p9.aes(color="z.abs() > 5"))
+ p9.labs(title="Chromium", y="concentration (µg/L)")
)
p2 = (
chromium
.query("abs(z) < 5")
>>
p9.ggplot(p9.aes(x="SampledDate", y="result", group="well_name"))
+ p9.geom_line(alpha=0.5)
+ p9.geom_point()
+ p9.labs(title="Chromium", y="concentration (µg/L)")
)
(p1 / p2)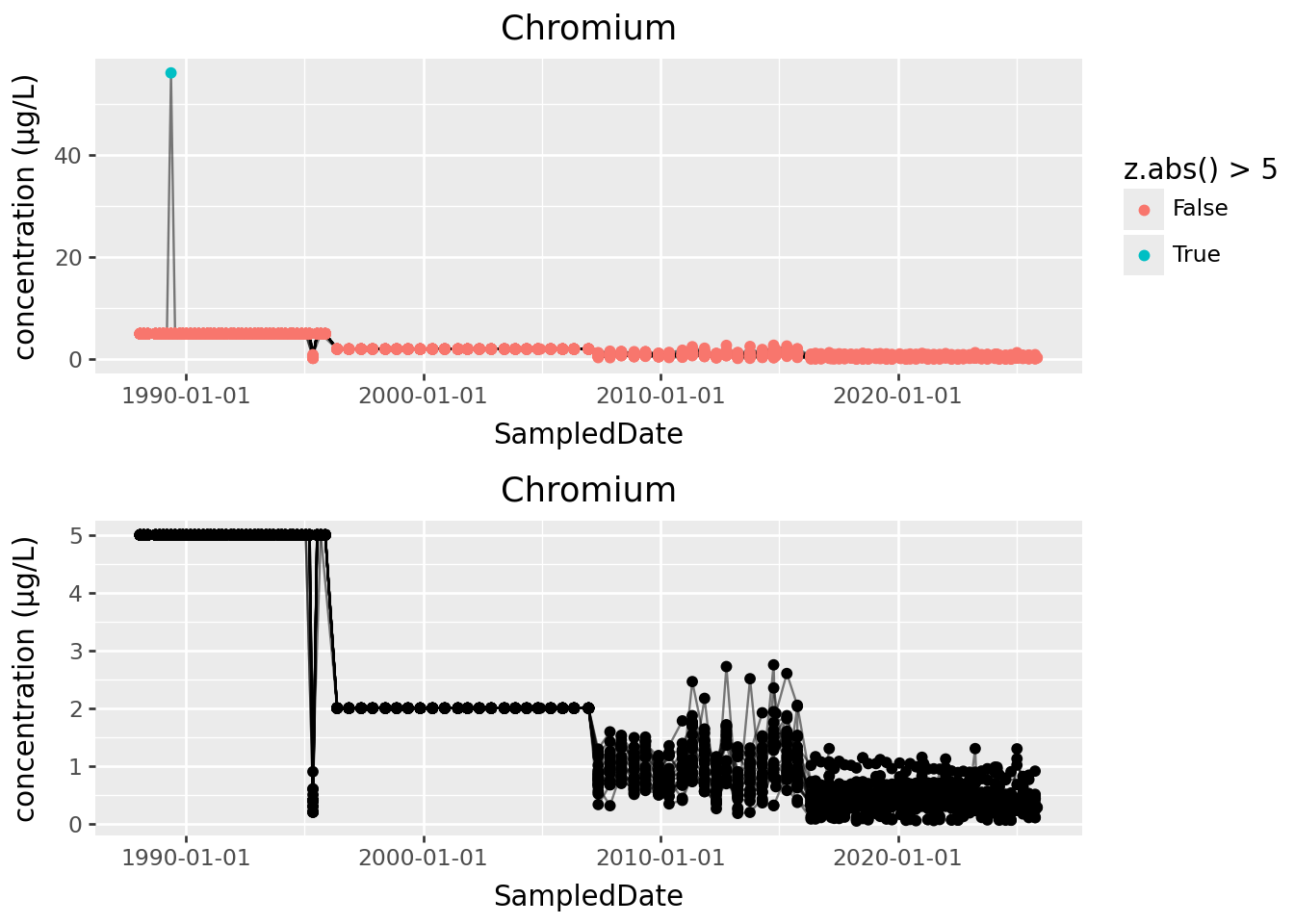
All the heavy metals
metals = ["Cadmium", "Chromium", "Copper", "Lead", "Nickel", "Zinc"]
(
gw
.loc[gw['ParameterName'].isin(metals), :]
>> p9.ggplot(p9.aes(x="SampledDate", y="result", color="well_orientation", group="well_name"))
+ p9.geom_line() + p9.geom_point(size=0.5, alpha=0.5)
+ p9.scale_y_log10()
+ p9.facet_grid(rows="ParameterName")
) + p9.theme(figure_size=(9,12))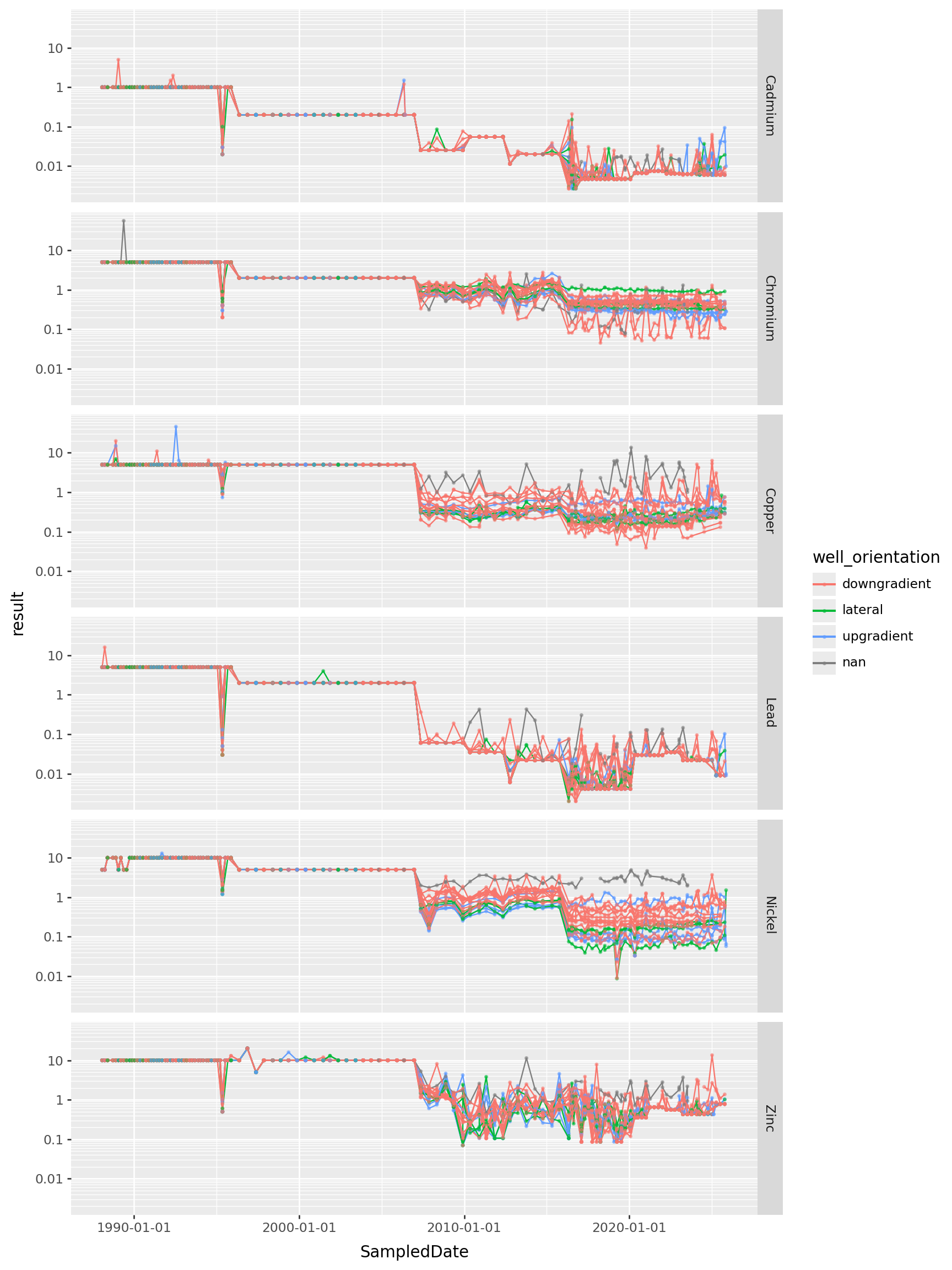
All the heavy metals, since 2016
(
gw
.loc[(gw['ParameterName'].isin(metals)) & (gw['SampledDate'].dt.year >= 2016), :]
>> p9.ggplot(p9.aes(x="SampledDate", y="result", color="well_orientation", group="well_name"))
+ p9.geom_line() + p9.geom_point(size=0.5, alpha=0.5)
+ p9.scale_y_log10()
+ p9.facet_grid(rows="ParameterName", scales="free_y")
) + p9.theme(figure_size=(9,12))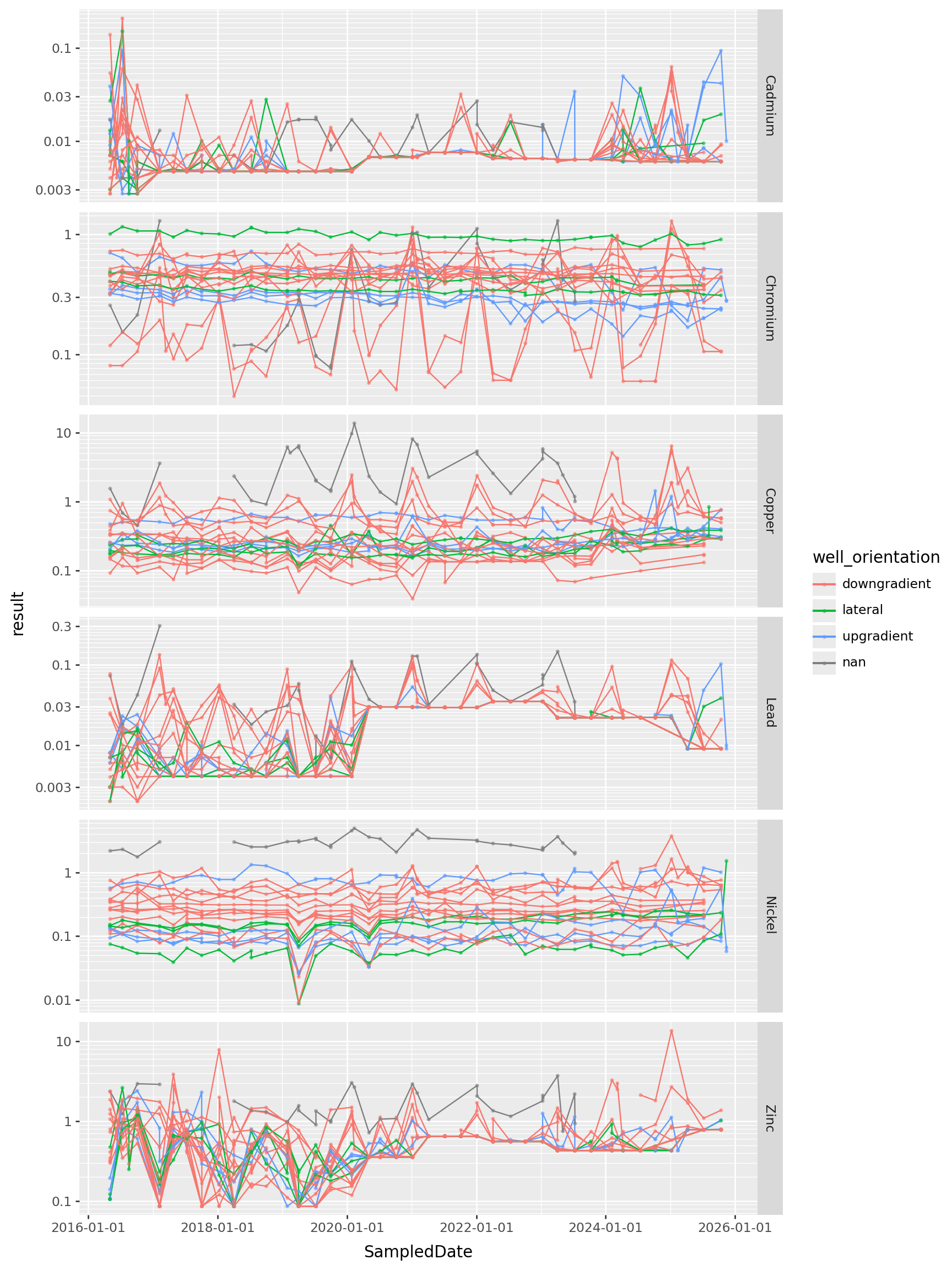
Takeaways?
Periodicity in water table elevation
Is the water table changing with the seasons? Looks like yes, although when it starts to come up again in the Fall seems to differ by year.
(
gw
.query("ParameterName == 'WaterTableElevation' and well_name == 'MW-17-1988'")
>>
p9.ggplot(p9.aes(x="SampledDate.dt.dayofyear", y="result", color='factor(SampledDate.dt.year)'))
+ p9.geom_point(size=0.5, alpha=0.75) + p9.geom_line()
+ p9.facet_wrap("well_name")
+ p9.labs(x='day of the year', y='level (ft)')
) + p9.theme(figure_size=(9,4))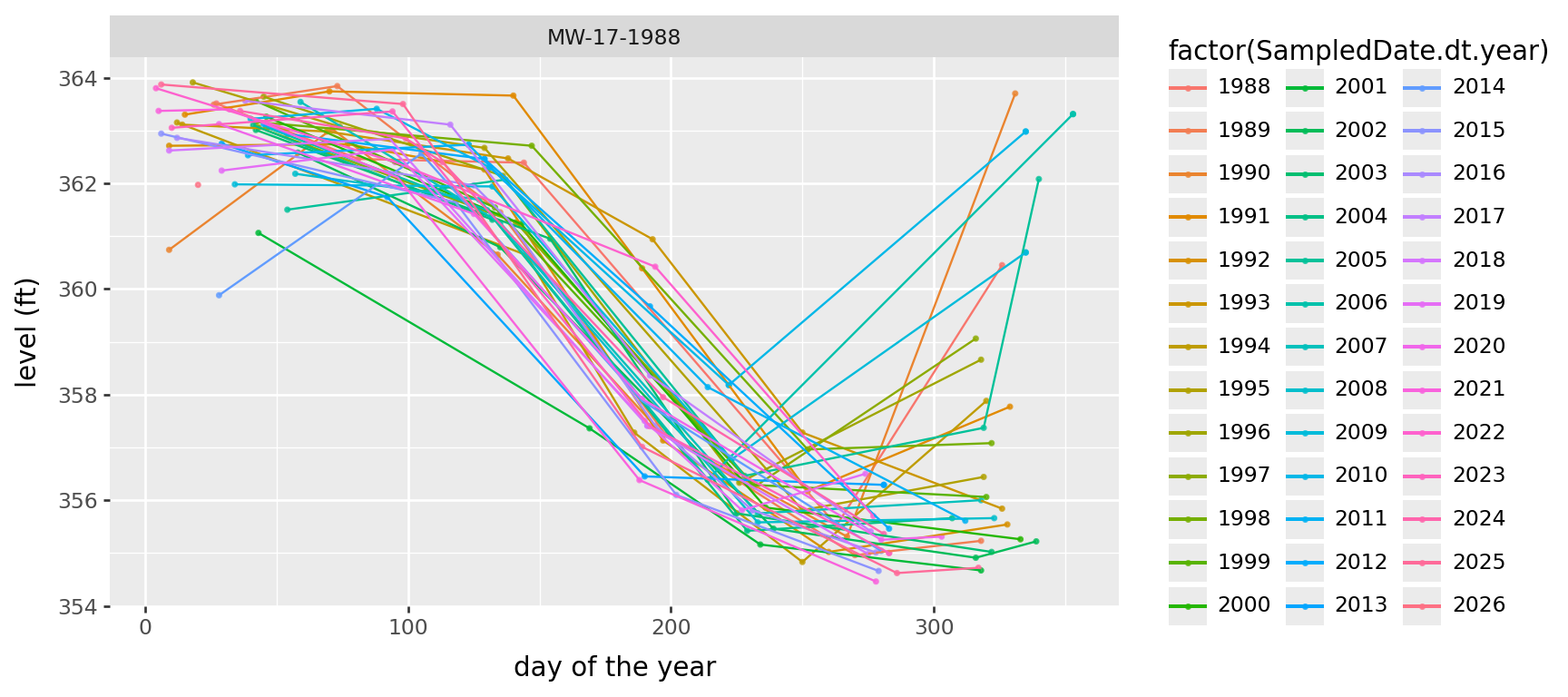
(
gw
.query("ParameterName == 'WaterTableElevation'")
>>
p9.ggplot(p9.aes(x="SampledDate.dt.dayofyear", y="result", color='factor(SampledDate.dt.year)'))
+ p9.geom_point(size=0.5, alpha=0.75) + p9.geom_line()
+ p9.facet_wrap("well_name")
+ p9.labs(x='day of the year', y='level (ft)')
) + p9.theme(figure_size=(9,15))/home/peter/micromamba/envs/ds435/lib/python3.13/site-packages/plotnine/layer.py:374: PlotnineWarning: geom_point : Removed 7 rows containing missing values.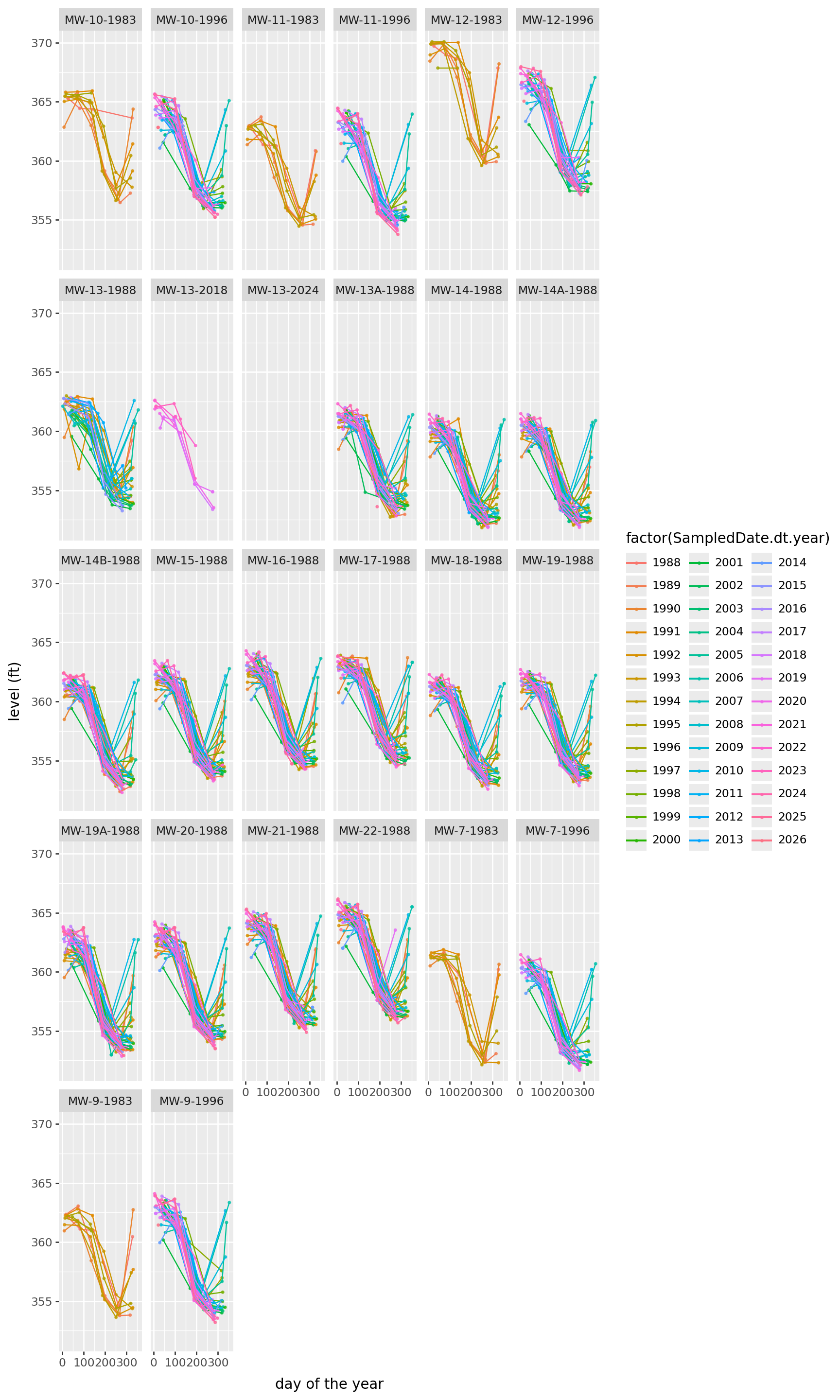
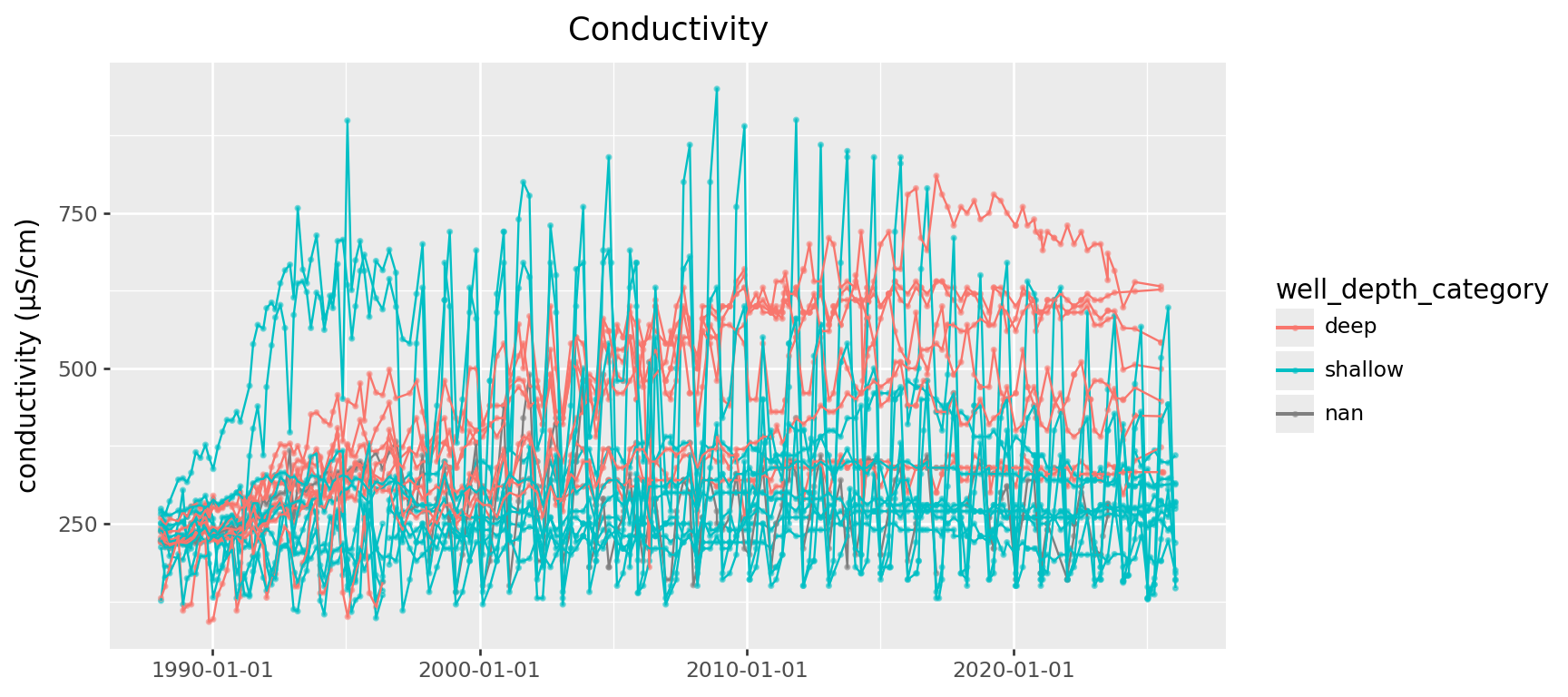
Conductivity
Let’s dive into conductivity:
Does the:
- location relative to the plant,
- or type of well
impact whether it’s:
- increasing/decreasing long term,
- or oscilating.
First step: separate out remove seasonal variation.
Model:
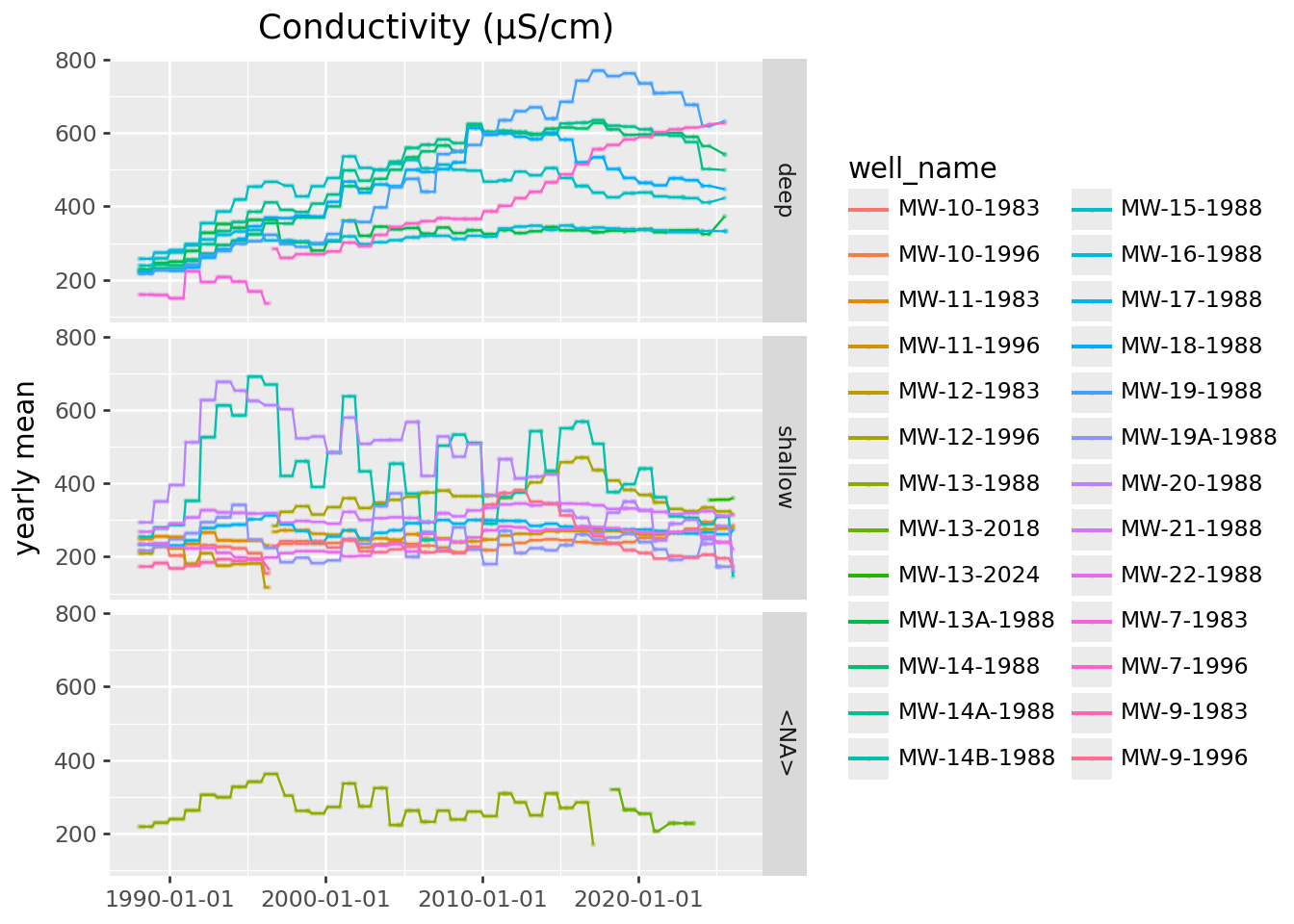
value = seasonal + trend(
conductivity
>>
p9.ggplot(p9.aes(x="SampledDate", y="mean_conductivity", color='well_depth_category', group='well_name'))
+ p9.geom_point(size=0.2, alpha=0.25) + p9.geom_line()
+ p9.labs(x='', y='yearly mean', title='Conductivity (µS/cm)')
) / (
conductivity
>>
p9.ggplot(p9.aes(x="SampledDate", y="diff_conductivity", color='well_depth_category', group='well_name'))
+ p9.geom_point(size=0.2, alpha=0.25) + p9.geom_line()
+ p9.labs(x='', y='seasonal difference')
) + p9.theme(figure_size=(9,6))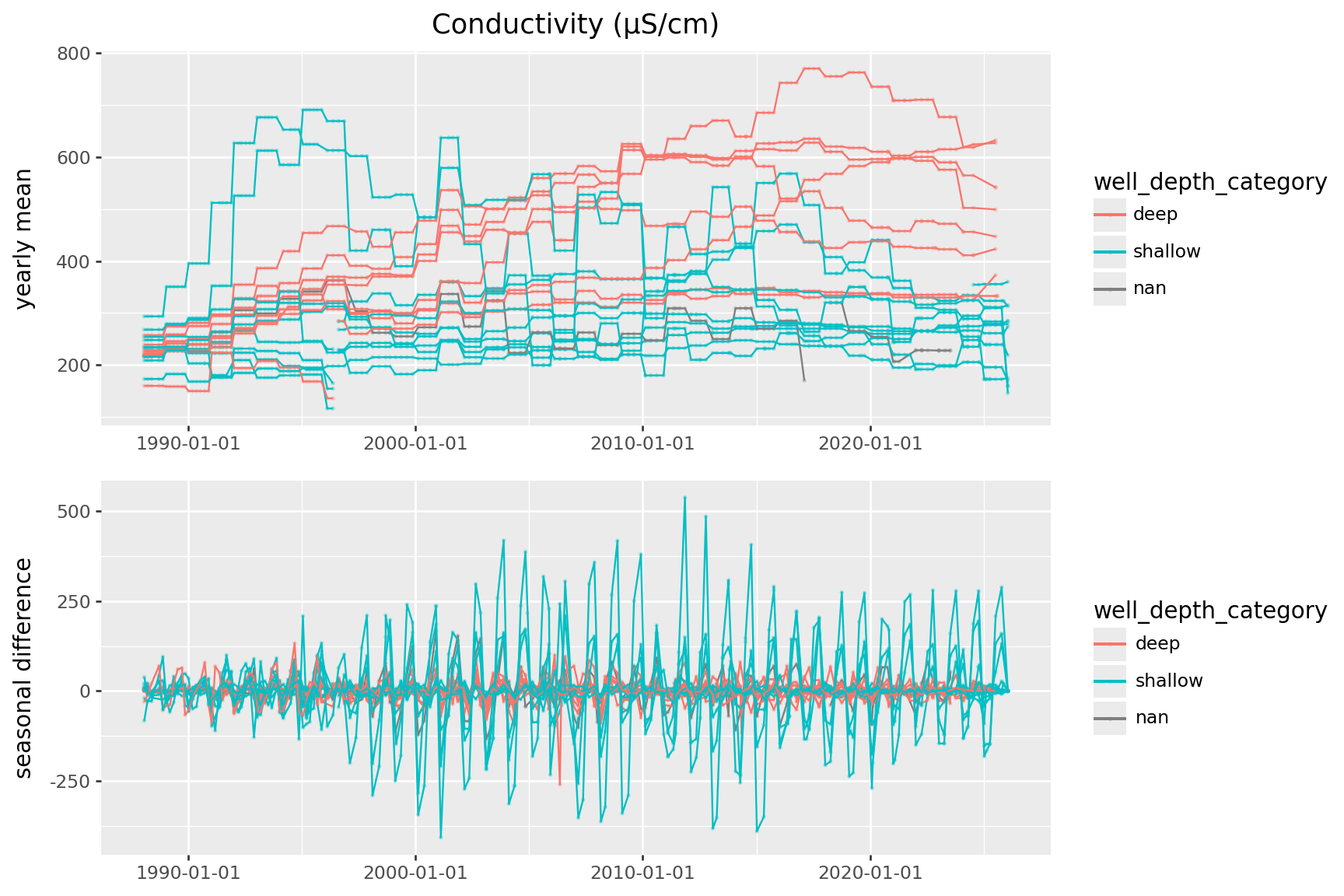
Per-well variability
The most variable wells are shallow:
| well_name | diff_conductivity | well_orientation | well_depth_category | |
|---|---|---|---|---|
| 3 | MW-11-1996 | 6.911335 | upgradient | shallow |
| 15 | MW-17-1988 | 7.788582 | lateral | shallow |
| 8 | MW-13-2024 | 9.622023 | downgradient | shallow |
| 14 | MW-16-1988 | 10.236792 | lateral | deep |
| 20 | MW-21-1988 | 11.955536 | lateral | shallow |
| 23 | MW-7-1996 | 12.942143 | downgradient | deep |
| 1 | MW-10-1996 | 13.141271 | upgradient | shallow |
| 21 | MW-22-1988 | 14.239555 | upgradient | shallow |
| 5 | MW-12-1996 | 15.75467 | upgradient | shallow |
| 10 | MW-14-1988 | 18.268072 | downgradient | deep |
| 25 | MW-9-1996 | 19.788127 | downgradient | shallow |
| 2 | MW-11-1983 | 21.207146 | upgradient | shallow |
| 11 | MW-14A-1988 | 23.100607 | downgradient | deep |
| 16 | MW-18-1988 | 24.80784 | downgradient | deep |
| 9 | MW-13A-1988 | 25.516837 | downgradient | deep |
| 24 | MW-9-1983 | 28.422397 | downgradient | shallow |
| 17 | MW-19-1988 | 31.878937 | downgradient | deep |
| 13 | MW-15-1988 | 36.719639 | downgradient | deep |
| 0 | MW-10-1983 | 39.299083 | upgradient | shallow |
| 7 | MW-13-2018 | 44.224813 | <NA> | <NA> |
| 22 | MW-7-1983 | 51.630195 | downgradient | deep |
| 4 | MW-12-1983 | 53.29468 | upgradient | shallow |
| 6 | MW-13-1988 | 56.293998 | <NA> | <NA> |
| 18 | MW-19A-1988 | 76.825087 | downgradient | shallow |
| 19 | MW-20-1988 | 109.612696 | downgradient | shallow |
| 12 | MW-14B-1988 | 201.358006 | downgradient | shallow |
Where are they?
And, mostly downgradient?
csdmap = make_colormap(csd.transform(np.log))
max_csd = csd.max()
legend_colors = { str(x) : csdmap(np.log(x)) for x in [1, int(max_csd/4), int(max_csd/2), int(max_csd*3/4), int(max_csd)] }
m = folium.Map(location=center, zoom_start=16)
for r in wells.itertuples():
add_marker(
m, r,
fill_color=csdmap(np.log(csd[r.well_name])),
)
add_legend(m, legend_colors, title='seasonality')
mLong-term trends:
Looks like the big increases in conductivity are all at deep wells.
Takeaways?
What’s correlated with what?
Question: what is “what”?
Refined question: which variables are correlated with each other, possibly considering seasonal variation or not?
Take 1: Per-observation correlation
| ParameterName | well_name | SampledDate | SampleNumber | Ammonia | Cadmium | ChemicalOxygenDemand | Chloride | Chromium | Conductivity | Copper | ... | Lead | Nickel | Nitrate | Temperature | TotalColiform | TotalDissolvedSolids | TotalOrganicCarbon | WaterTableElevation | Zinc | pH |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | MW-10-1983 | 1988-01-27 | 880068 | 0.1 | 1.0 | 5.0 | 1.9 | 5.0 | 238.0 | 5.0 | ... | 5.0 | 5.0 | 3.8 | <NA> | 1.0 | 181.0 | 2.0 | 365.28 | 10.0 | <NA> |
| 1 | MW-10-1983 | 1988-03-21 | 880248 | 0.1 | 1.0 | 10.0 | 4.5 | 5.0 | 238.0 | 5.0 | ... | 5.0 | 5.0 | 6.0 | <NA> | 1.0 | 196.0 | 2.0 | 364.44 | 10.0 | <NA> |
| 2 | MW-10-1983 | 1988-05-23 | 880426 | 0.1 | 1.0 | 10.0 | 3.1 | 5.0 | 198.0 | 5.0 | ... | 5.0 | 10.0 | 4.7 | <NA> | 9.0 | 151.0 | 2.0 | 364.38 | 10.0 | <NA> |
| 3 | MW-10-1983 | 1988-11-21 | 881128 | 0.1 | 1.0 | 10.0 | 2.8 | 5.0 | 192.0 | 15.0 | ... | 5.0 | 10.0 | 4.9 | <NA> | 70.0 | 164.0 | <NA> | 363.63 | 10.0 | <NA> |
| 4 | MW-10-1983 | 1989-01-26 | 890116 | 0.1 | 1.0 | 16.0 | 2.9 | 5.0 | 215.0 | 5.0 | ... | 5.0 | 5.0 | 4.0 | <NA> | 1.0 | 169.0 | <NA> | 365.3 | 10.0 | <NA> |
5 rows × 21 columns
Take 1: The correlation matrix
| ParameterName | Ammonia | Cadmium | ChemicalOxygenDemand | Chloride | Chromium | Conductivity | Copper | FecalColiform | Lead | Nickel | Nitrate | Temperature | TotalColiform | TotalDissolvedSolids | TotalOrganicCarbon | WaterTableElevation | Zinc | pH |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ParameterName | ||||||||||||||||||
| Ammonia | 1.000000 | -0.071792 | 0.160236 | 0.037876 | -0.074922 | 0.040336 | -0.099566 | -0.011911 | -0.081283 | -0.089971 | 0.019547 | -0.093406 | 0.029169 | 0.026352 | 0.034601 | -0.028478 | -0.101453 | -0.131407 |
| Cadmium | -0.071792 | 1.000000 | 0.281764 | -0.191022 | 0.863019 | -0.307538 | 0.710645 | 0.038601 | 0.960762 | 0.913679 | -0.084142 | -0.129080 | 0.096055 | -0.249209 | 0.507879 | 0.034189 | 0.749544 | 0.256278 |
| ChemicalOxygenDemand | 0.160236 | 0.281764 | 1.000000 | -0.115090 | 0.286894 | -0.158458 | 0.447788 | 0.076077 | 0.331570 | 0.373111 | -0.146574 | -0.142123 | 0.060277 | -0.157341 | 0.342511 | -0.000919 | 0.406129 | 0.038638 |
| Chloride | 0.037876 | -0.191022 | -0.115090 | 1.000000 | -0.161892 | 0.469068 | -0.207164 | -0.057571 | -0.204211 | -0.186110 | 0.285950 | 0.295613 | -0.048497 | 0.484853 | -0.211492 | -0.043363 | -0.195340 | 0.017063 |
| Chromium | -0.074922 | 0.863019 | 0.286894 | -0.161892 | 1.000000 | -0.259647 | 0.686521 | 0.032522 | 0.877213 | 0.839270 | -0.065953 | -0.133787 | 0.084074 | -0.205510 | 0.444899 | 0.014855 | 0.727691 | 0.391783 |
| Conductivity | 0.040336 | -0.307538 | -0.158458 | 0.469068 | -0.259647 | 1.000000 | -0.310711 | -0.088351 | -0.315467 | -0.281287 | 0.192999 | 0.372243 | -0.086641 | 0.964487 | -0.292122 | -0.329148 | -0.276213 | 0.164740 |
| Copper | -0.099566 | 0.710645 | 0.447788 | -0.207164 | 0.686521 | -0.310711 | 1.000000 | 0.079348 | 0.800688 | 0.827862 | -0.216518 | -0.249487 | 0.130510 | -0.263933 | 0.492412 | 0.000145 | 0.886070 | 0.090665 |
| FecalColiform | -0.011911 | 0.038601 | 0.076077 | -0.057571 | 0.032522 | -0.088351 | 0.079348 | 1.000000 | 0.049238 | 0.052259 | -0.050063 | -0.057543 | 0.574707 | -0.086840 | 0.211171 | 0.028351 | 0.040449 | -0.147483 |
| Lead | -0.081283 | 0.960762 | 0.331570 | -0.204211 | 0.877213 | -0.315467 | 0.800688 | 0.049238 | 1.000000 | 0.956844 | -0.117430 | -0.114198 | 0.099597 | -0.257265 | 0.505792 | 0.016446 | 0.850833 | 0.269656 |
| Nickel | -0.089971 | 0.913679 | 0.373111 | -0.186110 | 0.839270 | -0.281287 | 0.827862 | 0.052259 | 0.956844 | 1.000000 | -0.165641 | -0.194614 | 0.099505 | -0.228114 | 0.526614 | -0.014386 | 0.866621 | 0.162394 |
| Nitrate | 0.019547 | -0.084142 | -0.146574 | 0.285950 | -0.065953 | 0.192999 | -0.216518 | -0.050063 | -0.117430 | -0.165641 | 1.000000 | 0.211520 | -0.036594 | 0.257750 | -0.234865 | -0.049981 | -0.187701 | 0.201779 |
| Temperature | -0.093406 | -0.129080 | -0.142123 | 0.295613 | -0.133787 | 0.372243 | -0.249487 | -0.057543 | -0.114198 | -0.194614 | 0.211520 | 1.000000 | -0.216970 | 0.308916 | -0.324921 | -0.562670 | -0.131932 | 0.459840 |
| TotalColiform | 0.029169 | 0.096055 | 0.060277 | -0.048497 | 0.084074 | -0.086641 | 0.130510 | 0.574707 | 0.099597 | 0.099505 | -0.036594 | -0.216970 | 1.000000 | -0.087212 | 0.141671 | 0.017775 | 0.077223 | -0.146319 |
| TotalDissolvedSolids | 0.026352 | -0.249209 | -0.157341 | 0.484853 | -0.205510 | 0.964487 | -0.263933 | -0.086840 | -0.257265 | -0.228114 | 0.257750 | 0.308916 | -0.087212 | 1.000000 | -0.275284 | -0.325074 | -0.225765 | 0.156180 |
| TotalOrganicCarbon | 0.034601 | 0.507879 | 0.342511 | -0.211492 | 0.444899 | -0.292122 | 0.492412 | 0.211171 | 0.505792 | 0.526614 | -0.234865 | -0.324921 | 0.141671 | -0.275284 | 1.000000 | 0.063420 | 0.413437 | -0.406515 |
| WaterTableElevation | -0.028478 | 0.034189 | -0.000919 | -0.043363 | 0.014855 | -0.329148 | 0.000145 | 0.028351 | 0.016446 | -0.014386 | -0.049981 | -0.562670 | 0.017775 | -0.325074 | 0.063420 | 1.000000 | -0.022563 | -0.112165 |
| Zinc | -0.101453 | 0.749544 | 0.406129 | -0.195340 | 0.727691 | -0.276213 | 0.886070 | 0.040449 | 0.850833 | 0.866621 | -0.187701 | -0.131932 | 0.077223 | -0.225765 | 0.413437 | -0.022563 | 1.000000 | 0.262602 |
| pH | -0.131407 | 0.256278 | 0.038638 | 0.017063 | 0.391783 | 0.164740 | 0.090665 | -0.147483 | 0.269656 | 0.162394 | 0.201779 | 0.459840 | -0.146319 | 0.156180 | -0.406515 | -0.112165 | 0.262602 | 1.000000 |
Take 1: The correlation matrix, in a nice order:
| ParameterName | Lead | Nickel | Cadmium | Zinc | Copper | Chromium | TotalOrganicCarbon | Conductivity | ChemicalOxygenDemand | TotalDissolvedSolids | Chloride | Temperature | Nitrate | TotalColiform | FecalColiform | pH | WaterTableElevation | Ammonia |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ParameterName | ||||||||||||||||||
| Lead | 1.000000 | 0.956844 | 0.960762 | 0.850833 | 0.800688 | 0.877213 | 0.505792 | -0.315467 | 0.331570 | -0.257265 | -0.204211 | -0.114198 | -0.117430 | 0.099597 | 0.049238 | 0.269656 | 0.016446 | -0.081283 |
| Nickel | 0.956844 | 1.000000 | 0.913679 | 0.866621 | 0.827862 | 0.839270 | 0.526614 | -0.281287 | 0.373111 | -0.228114 | -0.186110 | -0.194614 | -0.165641 | 0.099505 | 0.052259 | 0.162394 | -0.014386 | -0.089971 |
| Cadmium | 0.960762 | 0.913679 | 1.000000 | 0.749544 | 0.710645 | 0.863019 | 0.507879 | -0.307538 | 0.281764 | -0.249209 | -0.191022 | -0.129080 | -0.084142 | 0.096055 | 0.038601 | 0.256278 | 0.034189 | -0.071792 |
| Zinc | 0.850833 | 0.866621 | 0.749544 | 1.000000 | 0.886070 | 0.727691 | 0.413437 | -0.276213 | 0.406129 | -0.225765 | -0.195340 | -0.131932 | -0.187701 | 0.077223 | 0.040449 | 0.262602 | -0.022563 | -0.101453 |
| Copper | 0.800688 | 0.827862 | 0.710645 | 0.886070 | 1.000000 | 0.686521 | 0.492412 | -0.310711 | 0.447788 | -0.263933 | -0.207164 | -0.249487 | -0.216518 | 0.130510 | 0.079348 | 0.090665 | 0.000145 | -0.099566 |
| Chromium | 0.877213 | 0.839270 | 0.863019 | 0.727691 | 0.686521 | 1.000000 | 0.444899 | -0.259647 | 0.286894 | -0.205510 | -0.161892 | -0.133787 | -0.065953 | 0.084074 | 0.032522 | 0.391783 | 0.014855 | -0.074922 |
| TotalOrganicCarbon | 0.505792 | 0.526614 | 0.507879 | 0.413437 | 0.492412 | 0.444899 | 1.000000 | -0.292122 | 0.342511 | -0.275284 | -0.211492 | -0.324921 | -0.234865 | 0.141671 | 0.211171 | -0.406515 | 0.063420 | 0.034601 |
| Conductivity | -0.315467 | -0.281287 | -0.307538 | -0.276213 | -0.310711 | -0.259647 | -0.292122 | 1.000000 | -0.158458 | 0.964487 | 0.469068 | 0.372243 | 0.192999 | -0.086641 | -0.088351 | 0.164740 | -0.329148 | 0.040336 |
| ChemicalOxygenDemand | 0.331570 | 0.373111 | 0.281764 | 0.406129 | 0.447788 | 0.286894 | 0.342511 | -0.158458 | 1.000000 | -0.157341 | -0.115090 | -0.142123 | -0.146574 | 0.060277 | 0.076077 | 0.038638 | -0.000919 | 0.160236 |
| TotalDissolvedSolids | -0.257265 | -0.228114 | -0.249209 | -0.225765 | -0.263933 | -0.205510 | -0.275284 | 0.964487 | -0.157341 | 1.000000 | 0.484853 | 0.308916 | 0.257750 | -0.087212 | -0.086840 | 0.156180 | -0.325074 | 0.026352 |
| Chloride | -0.204211 | -0.186110 | -0.191022 | -0.195340 | -0.207164 | -0.161892 | -0.211492 | 0.469068 | -0.115090 | 0.484853 | 1.000000 | 0.295613 | 0.285950 | -0.048497 | -0.057571 | 0.017063 | -0.043363 | 0.037876 |
| Temperature | -0.114198 | -0.194614 | -0.129080 | -0.131932 | -0.249487 | -0.133787 | -0.324921 | 0.372243 | -0.142123 | 0.308916 | 0.295613 | 1.000000 | 0.211520 | -0.216970 | -0.057543 | 0.459840 | -0.562670 | -0.093406 |
| Nitrate | -0.117430 | -0.165641 | -0.084142 | -0.187701 | -0.216518 | -0.065953 | -0.234865 | 0.192999 | -0.146574 | 0.257750 | 0.285950 | 0.211520 | 1.000000 | -0.036594 | -0.050063 | 0.201779 | -0.049981 | 0.019547 |
| TotalColiform | 0.099597 | 0.099505 | 0.096055 | 0.077223 | 0.130510 | 0.084074 | 0.141671 | -0.086641 | 0.060277 | -0.087212 | -0.048497 | -0.216970 | -0.036594 | 1.000000 | 0.574707 | -0.146319 | 0.017775 | 0.029169 |
| FecalColiform | 0.049238 | 0.052259 | 0.038601 | 0.040449 | 0.079348 | 0.032522 | 0.211171 | -0.088351 | 0.076077 | -0.086840 | -0.057571 | -0.057543 | -0.050063 | 0.574707 | 1.000000 | -0.147483 | 0.028351 | -0.011911 |
| pH | 0.269656 | 0.162394 | 0.256278 | 0.262602 | 0.090665 | 0.391783 | -0.406515 | 0.164740 | 0.038638 | 0.156180 | 0.017063 | 0.459840 | 0.201779 | -0.146319 | -0.147483 | 1.000000 | -0.112165 | -0.131407 |
| WaterTableElevation | 0.016446 | -0.014386 | 0.034189 | -0.022563 | 0.000145 | 0.014855 | 0.063420 | -0.329148 | -0.000919 | -0.325074 | -0.043363 | -0.562670 | -0.049981 | 0.017775 | 0.028351 | -0.112165 | 1.000000 | -0.028478 |
| Ammonia | -0.081283 | -0.089971 | -0.071792 | -0.101453 | -0.099566 | -0.074922 | 0.034601 | 0.040336 | 0.160236 | 0.026352 | 0.037876 | -0.093406 | 0.019547 | 0.029169 | -0.011911 | -0.131407 | -0.028478 | 1.000000 |
Take 2: Per-year (average) correlation:
yearly_gw = (
gw
.assign(year = lambda df: df['SampledDate'].dt.year)
.set_index(['well_name', 'ParameterName'])
.select_dtypes(("number", "datetime"))
.groupby(['well_name', 'year', 'ParameterName'])
.mean()
.reset_index()
.set_index(['well_name', 'SampledDate'])
.loc[:,('ParameterName', 'result')]
.pivot(columns='ParameterName', values='result')
.reset_index()
)
yearly_gw.head()| ParameterName | well_name | SampledDate | Ammonia | Cadmium | ChemicalOxygenDemand | Chloride | Chromium | Conductivity | Copper | FecalColiform | Lead | Nickel | Nitrate | Temperature | TotalColiform | TotalDissolvedSolids | TotalOrganicCarbon | WaterTableElevation | Zinc | pH |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | MW-10-1983 | 1988-03-24 00:00:00 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | 2.0 | <NA> | <NA> | <NA> |
| 1 | MW-10-1983 | 1988-05-23 12:00:00 | 0.1 | 1.0 | 8.75 | 3.075 | 5.0 | 216.5 | 7.5 | 10.75 | 5.0 | 7.5 | 4.85 | <NA> | 20.25 | 173.0 | <NA> | 364.4325 | 10.0 | <NA> |
| 2 | MW-10-1983 | 1989-05-29 19:12:00 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | 201.0 | <NA> | <NA> | <NA> | <NA> | <NA> |
| 3 | MW-10-1983 | 1989-06-18 20:00:00 | 0.1 | 1.0 | 11.0 | 2.183333 | 5.0 | 227.5 | 5.0 | 35.0 | 5.0 | 7.5 | 2.816667 | <NA> | <NA> | 171.5 | <NA> | 361.25 | 10.0 | <NA> |
| 4 | MW-10-1983 | 1989-07-17 14:24:00 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | 3.0 | <NA> | <NA> | <NA> |
Take 2: correlation matrix
Cor(temperature, ammonia) is NA, so we omit it:
| ParameterName | Nickel | Lead | Copper | Cadmium | Zinc | Chromium | Conductivity | TotalOrganicCarbon | TotalDissolvedSolids | ChemicalOxygenDemand | Chloride | pH | Nitrate | TotalColiform | FecalColiform | WaterTableElevation | Ammonia |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ParameterName | |||||||||||||||||
| Nickel | 1.000000 | 0.974307 | 0.873757 | 0.914269 | 0.839927 | 0.932369 | -0.443580 | 0.519879 | -0.329528 | 0.382999 | -0.258540 | -0.606327 | -0.262817 | 0.152710 | 0.113619 | 0.113584 | -0.111252 |
| Lead | 0.974307 | 1.000000 | 0.822724 | 0.963876 | 0.793517 | 0.962665 | -0.465164 | 0.459696 | -0.339230 | 0.329172 | -0.278216 | -0.354037 | -0.218795 | 0.153477 | 0.132039 | 0.102502 | -0.082002 |
| Copper | 0.873757 | 0.822724 | 1.000000 | 0.659877 | 0.947138 | 0.731038 | -0.486004 | 0.407220 | -0.390038 | 0.458215 | -0.293347 | -0.608708 | -0.268430 | 0.185043 | 0.146225 | 0.133345 | -0.145061 |
| Cadmium | 0.914269 | 0.963876 | 0.659877 | 1.000000 | 0.634862 | 0.960553 | -0.463534 | 0.466456 | -0.331818 | 0.237424 | -0.268653 | -0.177833 | -0.208729 | 0.149995 | 0.126472 | 0.099417 | -0.108828 |
| Zinc | 0.839927 | 0.793517 | 0.947138 | 0.634862 | 1.000000 | 0.707477 | -0.485474 | 0.366917 | -0.357321 | 0.440482 | -0.285125 | -0.351361 | -0.246009 | 0.152437 | 0.133209 | 0.111473 | -0.105533 |
| Chromium | 0.932369 | 0.962665 | 0.731038 | 0.960553 | 0.707477 | 1.000000 | -0.448873 | 0.453482 | -0.329363 | 0.288018 | -0.266127 | 0.334352 | -0.184797 | 0.143222 | 0.119331 | 0.083056 | -0.106301 |
| Conductivity | -0.443580 | -0.465164 | -0.486004 | -0.463534 | -0.485474 | -0.448873 | 1.000000 | -0.381327 | 0.983868 | -0.417007 | 0.493450 | 0.095067 | 0.173398 | -0.149138 | -0.142508 | -0.406708 | -0.024878 |
| TotalOrganicCarbon | 0.519879 | 0.459696 | 0.407220 | 0.466456 | 0.366917 | 0.453482 | -0.381327 | 1.000000 | -0.317408 | 0.361808 | -0.293275 | -0.498195 | -0.431739 | 0.215832 | 0.251730 | 0.168075 | 0.003362 |
| TotalDissolvedSolids | -0.329528 | -0.339230 | -0.390038 | -0.331818 | -0.357321 | -0.329363 | 0.983868 | -0.317408 | 1.000000 | -0.314850 | 0.450307 | 0.019255 | 0.251597 | -0.149055 | -0.148365 | -0.419909 | -0.007653 |
| ChemicalOxygenDemand | 0.382999 | 0.329172 | 0.458215 | 0.237424 | 0.440482 | 0.288018 | -0.417007 | 0.361808 | -0.314850 | 1.000000 | -0.254168 | -0.199517 | -0.280234 | 0.120198 | 0.121035 | 0.170892 | 0.376442 |
| Chloride | -0.258540 | -0.278216 | -0.293347 | -0.268653 | -0.285125 | -0.266127 | 0.493450 | -0.293275 | 0.450307 | -0.254168 | 1.000000 | -0.064295 | 0.274520 | -0.065459 | -0.082674 | 0.005607 | 0.029020 |
| pH | -0.606327 | -0.354037 | -0.608708 | -0.177833 | -0.351361 | 0.334352 | 0.095067 | -0.498195 | 0.019255 | -0.199517 | -0.064295 | 1.000000 | 0.199231 | -0.263690 | -0.222692 | -0.038003 | -0.036537 |
| Nitrate | -0.262817 | -0.218795 | -0.268430 | -0.208729 | -0.246009 | -0.184797 | 0.173398 | -0.431739 | 0.251597 | -0.280234 | 0.274520 | 0.199231 | 1.000000 | -0.062496 | -0.064400 | -0.031481 | 0.035847 |
| TotalColiform | 0.152710 | 0.153477 | 0.185043 | 0.149995 | 0.152437 | 0.143222 | -0.149138 | 0.215832 | -0.149055 | 0.120198 | -0.065459 | -0.263690 | -0.062496 | 1.000000 | 0.781211 | 0.111059 | 0.011857 |
| FecalColiform | 0.113619 | 0.132039 | 0.146225 | 0.126472 | 0.133209 | 0.119331 | -0.142508 | 0.251730 | -0.148365 | 0.121035 | -0.082674 | -0.222692 | -0.064400 | 0.781211 | 1.000000 | 0.045777 | -0.009722 |
| WaterTableElevation | 0.113584 | 0.102502 | 0.133345 | 0.099417 | 0.111473 | 0.083056 | -0.406708 | 0.168075 | -0.419909 | 0.170892 | 0.005607 | -0.038003 | -0.031481 | 0.111059 | 0.045777 | 1.000000 | 0.040060 |
| Ammonia | -0.111252 | -0.082002 | -0.145061 | -0.108828 | -0.105533 | -0.106301 | -0.024878 | 0.003362 | -0.007653 | 0.376442 | 0.029020 | -0.036537 | 0.035847 | 0.011857 | -0.009722 | 0.040060 | 1.000000 |
Take 3: Per-year averages, since 2016
recent_gw = (
gw
.query("SampledDate >= '01-01-2016'")
.assign(year = lambda df: df['SampledDate'].dt.year)
.set_index(['well_name', 'ParameterName'])
.select_dtypes(("number", "datetime"))
.groupby(['well_name', 'year', 'ParameterName'])
.mean()
.reset_index()
.set_index(['well_name', 'SampledDate'])
.loc[:,('ParameterName', 'result')]
.pivot(columns='ParameterName', values='result')
.reset_index()
)
recent_gw.head()| ParameterName | well_name | SampledDate | Ammonia | Cadmium | ChemicalOxygenDemand | Chloride | Chromium | Conductivity | Copper | FecalColiform | Lead | Nickel | Nitrate | Temperature | TotalColiform | TotalDissolvedSolids | TotalOrganicCarbon | WaterTableElevation | Zinc | pH |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 0 | MW-10-1996 | 2016-05-31 00:00:00 | 0.1 | <NA> | <NA> | 3.55 | <NA> | <NA> | <NA> | 1.0 | <NA> | <NA> | 3.525 | <NA> | 1.0 | 170.0 | <NA> | <NA> | <NA> | <NA> |
| 1 | MW-10-1996 | 2016-06-15 19:12:00 | <NA> | <NA> | <NA> | <NA> | <NA> | 240.0 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | 359.66 | <NA> | 7.02 |
| 2 | MW-10-1996 | 2016-07-16 16:00:00 | <NA> | <NA> | 5.0 | <NA> | 0.35 | <NA> | 0.265 | <NA> | 0.011667 | 0.1 | <NA> | <NA> | <NA> | <NA> | 0.5 | <NA> | 0.9192 | <NA> |
| 3 | MW-10-1996 | 2016-07-24 18:00:00 | <NA> | 0.028675 | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> | <NA> |
| 4 | MW-10-1996 | 2017-06-18 00:00:00 | 0.1 | 0.005083 | 5.0 | 3.516667 | 0.338 | 236.666667 | 0.236667 | 1.0 | 0.005233 | 0.085 | 3.816667 | <NA> | 1.0 | 166.666667 | 0.816667 | 360.96 | 0.746317 | 6.966667 |
Take 3: correlation matrix
| ParameterName | TotalOrganicCarbon | Copper | Nickel | Lead | Zinc | pH | Conductivity | TotalDissolvedSolids | ChemicalOxygenDemand | Cadmium | TotalColiform | Nitrate | FecalColiform | WaterTableElevation | Chloride | Ammonia | Chromium |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ParameterName | |||||||||||||||||
| TotalOrganicCarbon | 1.000000 | 0.890807 | 0.878169 | 0.753568 | 0.581503 | -0.498195 | -0.220552 | -0.251105 | 0.573421 | 0.283312 | 0.241063 | -0.378747 | 0.262582 | 0.216115 | -0.222661 | 0.056656 | 0.130319 |
| Copper | 0.890807 | 1.000000 | 0.909084 | 0.610292 | 0.628322 | -0.608708 | -0.322752 | -0.301071 | 0.352848 | 0.344804 | 0.310256 | -0.275047 | 0.167236 | 0.162293 | -0.073357 | -0.060376 | 0.135102 |
| Nickel | 0.878169 | 0.909084 | 1.000000 | 0.562719 | 0.650353 | -0.606327 | -0.201874 | -0.133684 | 0.311853 | 0.357334 | 0.215208 | -0.262286 | 0.359429 | 0.155727 | 0.074074 | -0.068902 | 0.055345 |
| Lead | 0.753568 | 0.610292 | 0.562719 | 1.000000 | 0.491133 | -0.354037 | -0.244525 | -0.213029 | 0.597012 | 0.283380 | 0.204370 | -0.205487 | 0.055351 | 0.257111 | -0.170673 | 0.128016 | 0.244156 |
| Zinc | 0.581503 | 0.628322 | 0.650353 | 0.491133 | 1.000000 | -0.351361 | -0.268892 | -0.204725 | 0.098982 | 0.432160 | 0.187908 | -0.213891 | 0.188980 | 0.084388 | -0.099996 | -0.117539 | 0.094888 |
| pH | -0.498195 | -0.608708 | -0.606327 | -0.354037 | -0.351361 | 1.000000 | 0.241367 | 0.190971 | -0.199517 | -0.177833 | -0.393534 | 0.289830 | -0.354504 | -0.231699 | 0.007443 | -0.031606 | 0.334352 |
| Conductivity | -0.220552 | -0.322752 | -0.201874 | -0.244525 | -0.268892 | 0.241367 | 1.000000 | 0.984912 | -0.186379 | -0.163036 | -0.123750 | 0.254431 | -0.110083 | -0.586880 | 0.369098 | 0.015135 | 0.125407 |
| TotalDissolvedSolids | -0.251105 | -0.301071 | -0.133684 | -0.213029 | -0.204725 | 0.190971 | 0.984912 | 1.000000 | -0.189485 | -0.240204 | -0.167343 | 0.325359 | -0.151245 | -0.622172 | 0.350088 | -0.028149 | 0.171769 |
| ChemicalOxygenDemand | 0.573421 | 0.352848 | 0.311853 | 0.597012 | 0.098982 | -0.199517 | -0.186379 | -0.189485 | 1.000000 | 0.119445 | -0.007333 | -0.196866 | 0.038757 | 0.224439 | -0.138664 | 0.602247 | 0.089381 |
| Cadmium | 0.283312 | 0.344804 | 0.357334 | 0.283380 | 0.432160 | -0.177833 | -0.163036 | -0.240204 | 0.119445 | 1.000000 | 0.244746 | -0.124114 | 0.324230 | 0.097763 | -0.202166 | -0.066868 | 0.078485 |
| TotalColiform | 0.241063 | 0.310256 | 0.215208 | 0.204370 | 0.187908 | -0.393534 | -0.123750 | -0.167343 | -0.007333 | 0.244746 | 1.000000 | -0.043623 | 0.983906 | 0.094847 | -0.107008 | -0.056771 | -0.083557 |
| Nitrate | -0.378747 | -0.275047 | -0.262286 | -0.205487 | -0.213891 | 0.289830 | 0.254431 | 0.325359 | -0.196866 | -0.124114 | -0.043623 | 1.000000 | -0.049502 | -0.214486 | 0.279763 | 0.018544 | 0.489560 |
| FecalColiform | 0.262582 | 0.167236 | 0.359429 | 0.055351 | 0.188980 | -0.354504 | -0.110083 | -0.151245 | 0.038757 | 0.324230 | 0.983906 | -0.049502 | 1.000000 | 0.095345 | -0.094697 | -0.023470 | -0.153142 |
| WaterTableElevation | 0.216115 | 0.162293 | 0.155727 | 0.257111 | 0.084388 | -0.231699 | -0.586880 | -0.622172 | 0.224439 | 0.097763 | 0.094847 | -0.214486 | 0.095345 | 1.000000 | 0.045799 | -0.030745 | -0.058456 |
| Chloride | -0.222661 | -0.073357 | 0.074074 | -0.170673 | -0.099996 | 0.007443 | 0.369098 | 0.350088 | -0.138664 | -0.202166 | -0.107008 | 0.279763 | -0.094697 | 0.045799 | 1.000000 | -0.012760 | 0.137604 |
| Ammonia | 0.056656 | -0.060376 | -0.068902 | 0.128016 | -0.117539 | -0.031606 | 0.015135 | -0.028149 | 0.602247 | -0.066868 | -0.056771 | 0.018544 | -0.023470 | -0.030745 | -0.012760 | 1.000000 | 0.080543 |
| Chromium | 0.130319 | 0.135102 | 0.055345 | 0.244156 | 0.094888 | 0.334352 | 0.125407 | 0.171769 | 0.089381 | 0.078485 | -0.083557 | 0.489560 | -0.153142 | -0.058456 | 0.137604 | 0.080543 | 1.000000 |
Takeaways:
Note: two variables are strongly correlated here if they tend to have larger-than-average values in the same year-well combinations.
So, this might be “the two variables are typically higher in the same wells”, or “the two variables are typically higher in the same years”, or some complicated combination.
“Look at the data”, part 3?
Next steps: brainstorm!
Ideas? Questions?
Do nearby wells get similar measurements?